MuDCoD: multi-subject community detection in personalized dynamic gene networks from single-cell RNA sequencing
- PMID: 37740957
- PMCID: PMC10564618
- DOI: 10.1093/bioinformatics/btad592
MuDCoD: multi-subject community detection in personalized dynamic gene networks from single-cell RNA sequencing
Abstract
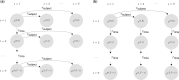
Motivation: With the wide availability of single-cell RNA-seq (scRNA-seq) technology, population-scale scRNA-seq datasets across multiple individuals and time points are emerging. While the initial investigations of these datasets tend to focus on standard analysis of clustering and differential expression, leveraging the power of scRNA-seq data at the personalized dynamic gene co-expression network level has the potential to unlock subject and/or time-specific network-level variation, which is critical for understanding phenotypic differences. Community detection from co-expression networks of multiple time points or conditions has been well-studied; however, none of the existing settings included networks from multiple subjects and multiple time points simultaneously. To address this, we develop Multi-subject Dynamic Community Detection (MuDCoD) for multi-subject community detection in personalized dynamic gene networks from scRNA-seq. MuDCoD builds on the spectral clustering framework and promotes information sharing among the networks of the subjects as well as networks at different time points. It clusters genes in the personalized dynamic gene networks and reveals gene communities that are variable or shared not only across time but also among subjects.
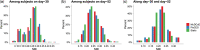
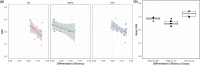
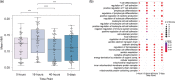
Results: Evaluation and benchmarking of MuDCoD against existing approaches reveal that MuDCoD effectively leverages apparent shared signals among networks of the subjects at individual time points, and performs robustly when there is no or little information sharing among the networks. Applications to population-scale scRNA-seq datasets of human-induced pluripotent stem cells during dopaminergic neuron differentiation and CD4+ T cell activation indicate that MuDCoD enables robust inference for identifying time-varying personalized gene modules. Our results illustrate how personalized dynamic community detection can aid in the exploration of subject-specific biological processes that vary across time.
Availability and implementation: MuDCoD is publicly available at https://github.com/bo1929/MuDCoD as a Python package. Implementation includes simulation and real-data experiments together with extensive documentation.
© The Author(s) 2023. Published by Oxford University Press.
Conflict of interest statement
None declared.
Figures


References
-
- Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B (Methodological) 1995;57:289–300.
-
- Chi Y, Song X, Zhou D et al. Evolutionary spectral clustering by incorporating temporal smoothness. In: Proceedings of the 13th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. New York, Association for Computing Machinery, 2007, 153–62.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

