Human neurotropic polyomavirus, JC virus, late coding region encodes a novel nuclear protein, ORF4, which targets the promyelocytic leukemia nuclear bodies (PML-NBs) and modulates their reorganization
- PMID: 37741199
- PMCID: PMC10602023
- DOI: 10.1016/j.virol.2023.109866
Human neurotropic polyomavirus, JC virus, late coding region encodes a novel nuclear protein, ORF4, which targets the promyelocytic leukemia nuclear bodies (PML-NBs) and modulates their reorganization
Abstract
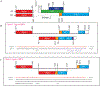
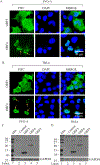
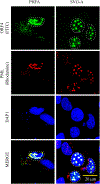
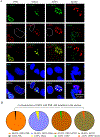
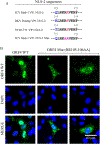
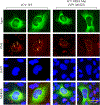
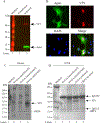
We previously reported the discovery and characterization of two novel proteins (ORF1 and ORF2) generated by the alternative splicing of the JC virus (JCV) late coding region. Here, we report the discovery and partial characterization of three additional novel ORFs from the same coding region, ORF3, ORF4 and ORF5, which potentially encode 70, 173 and 265 amino acid long proteins respectively. While ORF3 protein exhibits a uniform distribution pattern throughout the cells, we were unable to detect ORF5 expression. Surprisingly, ORF4 protein was determined to be the only JCV protein specifically targeting the promyelocytic leukemia nuclear bodies (PML-NBs) and inducing their reorganization in nucleus. Although ORF4 protein has a modest effect on JCV replication, it is implicated to play major roles during the JCV life cycle, perhaps by regulating the antiviral response of PML-NBs against JCV infections and thus facilitating the progression of the JCV-induced disease in infected individuals.
Keywords: BKV; DNA replication; Interferon; JCV, SV40; Merkel cell carcinoma virus; ORF, splicing; PML-NBs; Papillomavirus; Polyomavirus; Progressive multifocal leukoencephalopathy; Promyelocytic leukemia nuclear bodies; RNA splicing; Transcription.
Copyright © 2023 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of competing interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures





Similar articles
-
Discovery and characterization of novel trans-spliced products of human polyoma JC virus late transcripts from PML patients.J Cell Physiol. 2018 May;233(5):4137-4155. doi: 10.1002/jcp.26219. Epub 2017 Dec 18. J Cell Physiol. 2018. PMID: 29044559 Free PMC article.
-
Promyelocytic leukemia nuclear bodies provide a scaffold for human polyomavirus JC replication and are disrupted after development of viral inclusions in progressive multifocal leukoencephalopathy.J Neuropathol Exp Neurol. 2008 Apr;67(4):299-308. doi: 10.1097/NEN.0b013e31816a1dd3. J Neuropathol Exp Neurol. 2008. PMID: 18379438
-
[Development of intranuclear inclusions of human polyomavirus JC. Capsid proteins are assembled into virions at the PML nuclear bodies].Uirusu. 2006 Jun;56(1):17-25. doi: 10.2222/jsv.56.17. Uirusu. 2006. PMID: 17038808 Review. Japanese.
-
Functional reorganization of promyelocytic leukemia nuclear bodies during BK virus infection.mBio. 2011 Feb 8;2(1):e00281-10. doi: 10.1128/mBio.00281-10. Print 2011. mBio. 2011. PMID: 21304169 Free PMC article.
-
The Role of the JC Virus in Central Nervous System Tumorigenesis.Int J Mol Sci. 2020 Aug 28;21(17):6236. doi: 10.3390/ijms21176236. Int J Mol Sci. 2020. PMID: 32872288 Free PMC article. Review.
Cited by
-
Recent advances in discovery and functional analysis of the small proteins and microRNA expressed by polyomaviruses.Virology. 2025 Jan;602:110310. doi: 10.1016/j.virol.2024.110310. Epub 2024 Nov 22. Virology. 2025. PMID: 39612622 Free PMC article. Review.
References
-
- Becker KA, Florin L, Sapp C, Sapp M, 2003. Dissection of human papillomavirus type 33 L2 domains involved in nuclear domains (ND) 10 homing and reorganization. Virology 314, 161–167. - PubMed
-
- Berger JR, 1992. PML in AIDS. Neurology 42, 1845–1846. - PubMed
-
- Berger JR, 2000. Progressive Multifocal Leukoencephalopathy. Curr Treat Options Neurol 2, 361–368. - PubMed
-
- Berger JR, 2007. Progressive multifocal leukoencephalopathy. Curr Neurol Neurosci Rep 7, 461–469. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

