This is a preprint.
Reentrant DNA shells tune polyphosphate condensate size
- PMID: 37745474
- PMCID: PMC10515899
- DOI: 10.1101/2023.09.13.557044
Reentrant DNA shells tune polyphosphate condensate size
Update in
-
Reentrant DNA shells tune polyphosphate condensate size.Nat Commun. 2024 Oct 26;15(1):9258. doi: 10.1038/s41467-024-53469-x. Nat Commun. 2024. PMID: 39462120 Free PMC article.
Abstract
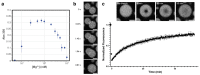
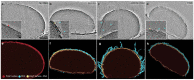
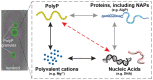
The ancient, inorganic biopolymer polyphosphate (polyP) occurs in all three domains of life and affects myriad cellular processes. An intriguing feature of polyP is its frequent proximity to chromatin, and in the case of many bacteria, its occurrence in the form of magnesium-enriched condensates embedded in the nucleoid, particularly in response to stress. The physical basis of the interaction between polyP and DNA, two fundamental anionic biopolymers, and the resulting effects on the organization of both the nucleoid and polyP condensates remain poorly understood. Given the essential role of magnesium ions in the coordination of polymeric phosphate species, we hypothesized that a minimal system of polyP, magnesium ions, and DNA (polyP-Mg2+-DNA) would capture key features of the interplay between the condensates and bacterial chromatin. We find that DNA can profoundly affect polyP-Mg2+ coacervation even at concentrations several orders of magnitude lower than found in the cell. The DNA forms shells around polyP-Mg2+ condensates and these shells show reentrant behavior, primarily forming in the concentration range close to polyP-Mg2+ charge neutralization. This surface association tunes both condensate size and DNA morphology in a manner dependent on DNA properties, including length and concentration. Our work identifies three components that could form the basis of a central and tunable interaction hub that interfaces with cellular interactors. These studies will inform future efforts to understand the basis of polyP granule composition and consolidation, as well as the potential capacity of these mesoscale assemblies to remodel chromatin in response to diverse stressors at different length and time scales.
Figures


References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
