This is a preprint.
Genetic Variants That Modify the Neuroendocrine Regulation of Foraging Behavior in C. elegans
- PMID: 37745484
- PMCID: PMC10515746
- DOI: 10.1101/2023.09.09.556976
Genetic Variants That Modify the Neuroendocrine Regulation of Foraging Behavior in C. elegans
Update in
-
Genetic variants that modify neuroendocrine gene expression and foraging behavior of C. elegans.Sci Adv. 2024 Jun 14;10(24):eadk9481. doi: 10.1126/sciadv.adk9481. Epub 2024 Jun 12. Sci Adv. 2024. PMID: 38865452 Free PMC article.
Abstract
The molecular mechanisms underlying diversity in animal behavior are not well understood. A major experimental challenge is determining the contribution of genetic variants that affect neuronal gene expression to differences in behavioral traits. The neuroendocrine TGF-beta ligand, DAF-7, regulates diverse behavioral responses of Caenorhabditis elegans to bacterial food and pathogens. The dynamic neuron-specific expression of daf-7 is modulated by environmental and endogenous bacteria-derived cues. Here, we investigated natural variation in the expression of daf-7 from the ASJ pair of chemosensory neurons and identified common variants in gap-2, encoding a GTPase-Activating Protein homologous to mammalian SynGAP proteins, which modify daf-7 expression cell-non-autonomously and promote exploratory foraging behavior in a DAF-7-dependent manner. Our data connect natural variation in neuron-specific gene expression to differences in behavior and suggest that genetic variation in neuroendocrine signaling pathways mediating host-microbe interactions may give rise to diversity in animal behavior.
Conflict of interest statement
Competing interests: Authors declare that they have no competing interests.
Figures




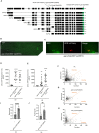
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
