The Penicillium chrysogenum tom1 Gene a Major Target of Transcription Factor MAT1-1-1 Encodes a Nuclear Protein Involved in Sporulation
- PMID: 37746180
- PMCID: PMC10512297
- DOI: 10.3389/ffunb.2022.937023
The Penicillium chrysogenum tom1 Gene a Major Target of Transcription Factor MAT1-1-1 Encodes a Nuclear Protein Involved in Sporulation
Abstract
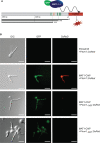
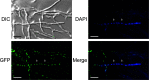
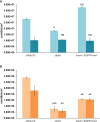
Fungal mating-type loci (MAT) encode transcription factors (TFs) MAT1-1-1 and MAT1-2-1, which govern sexual reproduction as well as other developmental processes. In Penicillium chrysogenum, the major producer of the beta-lactam antibiotic penicillin, a recent chromatin immunoprecipitation followed by sequencing (ChIP-seq) analysis identified 254 genes as direct targets of MAT1-1-1, many of which encode thus far uncharacterized proteins. Here, we characterized one of the major targets of MAT1-1-1, the tom1 gene, which encodes a protein highly conserved within the group of Eurotiomycetes fungi. Using fluorescence microscopy, we demonstrated binding of MAT1-1-1 to the tom1 promoter by reporter gene analysis. Extensive electrophoretic mobility shift assays (EMSAs) further showed that the promoter sequence of tom1 is bound in vitro by both MAT1-1-1 and MAT1-2-1. This indicated an interaction between the two TFs, which was verified by yeast two-hybrid analysis. The sequence of tom1 carries a nuclear localization sequence, and indeed its nuclear localization was verified by fluorescence microscopy. The in vivo function of tom1 was investigated using tom1 deletion strains, as well as a complementing strain where the wild-type tom1 gene was reintroduced. We found a clear sporulation defect in the deletion strain, which became more evident when the fungi were grown at an elevated temperature of 31°C.
Keywords: Mating-type transcription factor; Penicillium chrysogenum; electrophoretic mobility shift assay; fluorescence microscopy; sporulation; yeast two-hybrid analysis.
Copyright © 2022 Ramšak and Kück.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
A MAT1-2 wild-type strain from Penicillium chrysogenum: functional mating-type locus characterization, genome sequencing and mating with an industrial penicillin-producing strain.Mol Microbiol. 2015 Mar;95(5):859-74. doi: 10.1111/mmi.12909. Epub 2015 Jan 30. Mol Microbiol. 2015. PMID: 25521009 Free PMC article.
-
Genome-wide identification of target genes of a mating-type α-domain transcription factor reveals functions beyond sexual development.Mol Microbiol. 2015 Jun;96(5):1002-22. doi: 10.1111/mmi.12987. Epub 2015 Mar 28. Mol Microbiol. 2015. PMID: 25728030
-
Molecular analysis of mating type loci from the mycophenolic acid producer Penicillium brevicompactum: Phylogeny and MAT protein characterization suggest a cryptic sexual life cycle.Fungal Biol. 2020 Sep;124(9):821-833. doi: 10.1016/j.funbio.2020.07.006. Epub 2020 Jul 29. Fungal Biol. 2020. PMID: 32883432
-
Key role of LaeA and velvet complex proteins on expression of β-lactam and PR-toxin genes in Penicillium chrysogenum: cross-talk regulation of secondary metabolite pathways.J Ind Microbiol Biotechnol. 2017 May;44(4-5):525-535. doi: 10.1007/s10295-016-1830-y. Epub 2016 Aug 26. J Ind Microbiol Biotechnol. 2017. PMID: 27565675 Review.
-
[Classification of dermatophytes by mating type (MAT) gene analysis].Med Mycol J. 2012;53(3):175-8. doi: 10.3314/mmj.53.175. Med Mycol J. 2012. PMID: 23149351 Review. Japanese.
Cited by
-
Three-Dimensional Structural Heteromorphs of Mating-Type Proteins in Hirsutella sinensis and the Natural Cordyceps sinensis Insect-Fungal Complex.J Fungi (Basel). 2025 Mar 23;11(4):244. doi: 10.3390/jof11040244. J Fungi (Basel). 2025. PMID: 40278065 Free PMC article.
References
-
- Badgett T. C., Staben C. (1999). Interaction Between and Transactivation by the Mating Type Polypeptides of Neurospora crassa . Fungal Genet. Newsl. 46S, 127.
LinkOut - more resources
Full Text Sources

