Dissecting the immune discrepancies in mouse liver allograft tolerance and heart/kidney allograft rejection
- PMID: 37748771
- PMCID: PMC10905343
- DOI: 10.1111/cpr.13555
Dissecting the immune discrepancies in mouse liver allograft tolerance and heart/kidney allograft rejection
Abstract
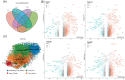
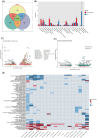
The liver is the most tolerogenic of transplanted organs. However, the mechanisms underlying liver transplant tolerance are not well understood. The comparison between liver transplantation tolerance and heart/kidney transplantation rejection will deepen our understanding of tolerance and rejection in solid organs. Here, we built a mouse model of liver, heart and kidney allograft and performed single-cell RNA sequencing of 66,393 cells to describe the cell composition and immune cell interactions at the early stage of tolerance or rejection. We also performed bulk RNA-seq of mouse liver allografts from Day 7 to Day 60 post-transplantation to map the dynamic transcriptional variation in spontaneous tolerance. The transcriptome of lymphocytes and myeloid cells were characterized and compared in three types of organ allografts. Cell-cell interaction networks reveal the coordinated function of Kupffer cells, macrophages and their associated metabolic processes, including insulin receptor signalling and oxidative phosphorylation in tolerance induction. Cd11b+ dendritic cells (DCs) in liver allografts were found to inhibit cytotoxic T cells by secreting anti-inflammatory cytokines such as Il10. In summary, we profiled single-cell transcriptome analysis of mouse solid organ allografts. We characterized the immune microenvironment of mouse organ allografts in the acute rejection state (heart, kidney) and tolerance state (liver).
© 2023 The Authors. Cell Proliferation published by Beijing Institute for Stem Cell and Regenerative Medicine and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no competing interests.
Figures



References
-
- Loupy A, Lefaucheur C. Antibody‐mediated rejection of solid‐organ allografts. New Engl J Med. 2018;379(12):1150‐1160. - PubMed
-
- Thomson AW, Vionnet J, Sanchez‐Fueyo A. Understanding, predicting and achieving liver transplant tolerance: from bench to bedside. Nat Rev Gastroentrol Hepatol. 2020;17(12):719‐739. - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

