Discovery of Nedd4 auto-ubiquitination inhibitors
- PMID: 37749144
- PMCID: PMC10520017
- DOI: 10.1038/s41598-023-42997-z
Discovery of Nedd4 auto-ubiquitination inhibitors
Abstract
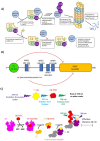
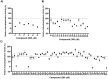
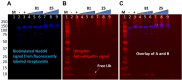
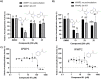
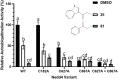
E3 ubiquitin ligases are critical to the protein degradation pathway by catalyzing the final step in protein ubiquitination by mediating ubiquitin transfer from E2 enzymes to target proteins. Nedd4 is a HECT domain-containing E3 ubiquitin ligase with a wide range of protein targets, the dysregulation of which has been implicated in myriad pathologies, including cancer and Parkinson's disease. Towards the discovery of compounds disrupting the auto-ubiquitination activity of Nedd4, we developed and optimized a TR-FRET assay for high-throughput screening. Through selective screening of a library of potentially covalent compounds, compounds 25 and 81 demonstrated apparent IC50 values of 52 µM and 31 µM, respectively. Tandem mass spectrometry (MS/MS) analysis confirmed that 25 and 81 were covalently bound to Nedd4 cysteine residues (Cys182 and Cys867). In addition, 81 also adducted to Cys627. Auto-ubiquitination assays of Nedd4 mutants featuring alanine substitutions for each of these cysteines suggested that the mode of inhibition of these compounds occurs through blocking the catalytic Cys867. The discovery of these inhibitors could enable the development of therapeutics for various diseases caused by Nedd4 E3 ligase dysregulation.
© 2023. Springer Nature Limited.
Conflict of interest statement
The authors declare no competing interests.
Figures




Similar articles
-
Comparative analysis of the catalytic regulation of NEDD4-1 and WWP2 ubiquitin ligases.J Biol Chem. 2019 Nov 15;294(46):17421-17436. doi: 10.1074/jbc.RA119.009211. Epub 2019 Oct 2. J Biol Chem. 2019. PMID: 31578285 Free PMC article.
-
Calcium activates Nedd4 E3 ubiquitin ligases by releasing the C2 domain-mediated auto-inhibition.J Biol Chem. 2010 Apr 16;285(16):12279-88. doi: 10.1074/jbc.M109.086405. Epub 2010 Feb 19. J Biol Chem. 2010. PMID: 20172859 Free PMC article.
-
Regulation of Nedd4-2 self-ubiquitination and stability by a PY motif located within its HECT-domain.Biochem J. 2008 Oct 1;415(1):155-63. doi: 10.1042/BJ20071708. Biochem J. 2008. PMID: 18498246
-
The role of NEDD4 related HECT-type E3 ubiquitin ligases in defective autophagy in cancer cells: molecular mechanisms and therapeutic perspectives.Mol Med. 2023 Mar 14;29(1):34. doi: 10.1186/s10020-023-00628-3. Mol Med. 2023. PMID: 36918822 Free PMC article. Review.
-
Emerging roles of the HECT E3 ubiquitin ligases in gastric cancer.Pathol Oncol Res. 2023 Feb 7;29:1610931. doi: 10.3389/pore.2023.1610931. eCollection 2023. Pathol Oncol Res. 2023. PMID: 36825281 Free PMC article. Review.
Cited by
-
Therapeutic Implications and Regulations of Protein Post-translational Modifications in Parkinsons Disease.Cell Mol Neurobiol. 2024 Jul 3;44(1):53. doi: 10.1007/s10571-024-01471-8. Cell Mol Neurobiol. 2024. PMID: 38960968 Free PMC article. Review.
-
Ubiquitination Enzymes in Cancer, Cancer Immune Evasion, and Potential Therapeutic Opportunities.Cells. 2025 Jan 7;14(2):69. doi: 10.3390/cells14020069. Cells. 2025. PMID: 39851497 Free PMC article. Review.
-
LZTR1 is a melanoma oncogene that promotes invasion and suppresses apoptosis.Oncogene. 2025 Aug 30. doi: 10.1038/s41388-025-03538-2. Online ahead of print. Oncogene. 2025. PMID: 40885854
-
Cardiotoxicity of Chemotherapy: A Multi-OMIC Perspective.J Xenobiot. 2025 Jan 8;15(1):9. doi: 10.3390/jox15010009. J Xenobiot. 2025. PMID: 39846541 Free PMC article. Review.
-
Structure-based design of potent and selective inhibitors of the HECT ligase NEDD4.Commun Chem. 2025 May 28;8(1):164. doi: 10.1038/s42004-025-01557-4. Commun Chem. 2025. PMID: 40437006 Free PMC article.
References
-
- Akutsu M, Dikic I, Bremm A. Ubiquitin chain diversity at a glance. J. Cell Sci. 2016;129:875–880. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

