IL-12 sensing in neurons induces neuroprotective CNS tissue adaptation and attenuates neuroinflammation in mice
- PMID: 37749256
- PMCID: PMC10545539
- DOI: 10.1038/s41593-023-01435-z
IL-12 sensing in neurons induces neuroprotective CNS tissue adaptation and attenuates neuroinflammation in mice
Abstract
Interleukin-12 (IL-12) is a potent driver of type 1 immunity. Paradoxically, in autoimmune conditions, including of the CNS, IL-12 reduces inflammation. The underlying mechanism behind these opposing properties and the involved cellular players remain elusive. Here we map IL-12 receptor (IL-12R) expression to NK and T cells as well as neurons and oligodendrocytes. Conditionally ablating the IL-12R across these cell types in adult mice and assessing their susceptibility to experimental autoimmune encephalomyelitis revealed that the neuroprotective role of IL-12 is mediated by neuroectoderm-derived cells, specifically neurons, and not immune cells. In human brain tissue from donors with multiple sclerosis, we observe an IL-12R distribution comparable to mice, suggesting similar mechanisms in mice and humans. Combining flow cytometry, bulk and single-nucleus RNA sequencing, we reveal an IL-12-induced neuroprotective tissue adaption preventing early neurodegeneration and sustaining trophic factor release during neuroinflammation, thereby maintaining CNS integrity in mice.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures










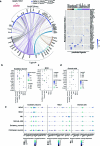


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

