Cbx7 promotes the generation of induced pluripotent stem cells
- PMID: 37753387
- PMCID: PMC10518684
- DOI: 10.1016/j.reth.2023.09.008
Cbx7 promotes the generation of induced pluripotent stem cells
Abstract
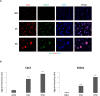
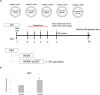
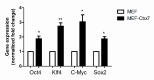
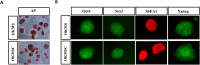
The iPS cells were discovered in 2006. With their ability to differentiate into cells of all three germ layers, iPS cells have great potential for clinical applications. Oct4, Sox2, c-Myc, and Klf4 were identified as the most effective factors for generating iPS cells. Despite this, iPS cells manufactured with these factors would still be inefficient. As a member of the chromobox family, chromobox protein homolog 7 (Cbx7) binds to PRC1 and PRC2 to inhibit genes involved in differentiation. A decrease in the expression of Cbx7 is observed during embryonic stem cell differentiation. Currently, no report discusses the role of Cbx7 in the production of iPS cells. In this study, we hypothesized that Cbx7 could increase iPS cell generation. We confirmed that Cbx7 is highly expressed in pluripotent stem cells (including ES and iPS cells). In addition, transfecting Cbx7 into fibroblasts increased Oct4, Sox2, c-Myc, and Klf4 expression. Moreover, we describe a novel approach to producing iPS cells using Cbx7 in combination with Oct4, Sox2, c-Myc, and Klf4. In summary, we have demonstrated that Cbx7 enhances the reprogramming of iPS cells and characterized the stemness and pluripotency of iPS cells.
Keywords: Cbx7; Induced pluripotent stem cells; Yamanaka factors.
© 2023 The Japanese Society for Regenerative Medicine. Production and hosting by Elsevier B.V.
Conflict of interest statement
The authors have no conflicts of interest to declare.
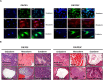
Figures

Similar articles
-
Murine pluripotent stem cells that escape differentiation inside teratomas maintain pluripotency.PeerJ. 2018 Jan 4;6:e4177. doi: 10.7717/peerj.4177. eCollection 2018. PeerJ. 2018. PMID: 29312817 Free PMC article.
-
Reprogramming of adult human testicular cells by four transcription factors (OCT4, SOX2, KLF4, and C-MYC).Reprod Med Biol. 2011 Feb 16;10(2):105-112. doi: 10.1007/s12522-011-0077-6. eCollection 2011 Jun. Reprod Med Biol. 2011. PMID: 29699086 Free PMC article.
-
Trib2 regulates the pluripotency of embryonic stem cells and enhances reprogramming efficiency.Exp Mol Med. 2017 Nov 24;49(11):e401. doi: 10.1038/emm.2017.191. Exp Mol Med. 2017. PMID: 29170476 Free PMC article.
-
Embryonic stem cell microRNAs: defining factors in induced pluripotent (iPS) and cancer (CSC) stem cells?Curr Stem Cell Res Ther. 2009 Sep;4(3):168-77. doi: 10.2174/157488809789057400. Curr Stem Cell Res Ther. 2009. PMID: 19492978 Review.
-
Generation of clinically relevant "induced pluripotent stem" (iPS) cells.J Stem Cells. 2011;6(3):109-27. J Stem Cells. 2011. PMID: 23264997 Review.
Cited by
-
Biomimetic Scaffolds-A Novel Approach to Three Dimensional Cell Culture Techniques for Potential Implementation in Tissue Engineering.Nanomaterials (Basel). 2024 Mar 16;14(6):531. doi: 10.3390/nano14060531. Nanomaterials (Basel). 2024. PMID: 38535679 Free PMC article. Review.
-
Chromobox protein homolog 7 suppresses the stem-like phenotype of glioblastoma cells by regulating the myosin heavy chain 9-NF-κB signaling pathway.Cell Death Discov. 2025 Feb 23;11(1):74. doi: 10.1038/s41420-025-02362-7. Cell Death Discov. 2025. PMID: 39988672 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Research Materials

