Comprehensive Genome and Transcriptome Analysis Identifies SLCO3A1 Associated with Aggressive Behavior in Pigs
- PMID: 37759782
- PMCID: PMC10526945
- DOI: 10.3390/biom13091381
Comprehensive Genome and Transcriptome Analysis Identifies SLCO3A1 Associated with Aggressive Behavior in Pigs
Abstract
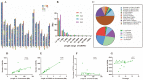
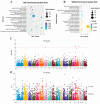
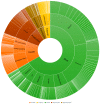
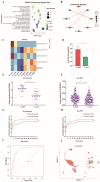
Copy number variation (CNV) represents a significant reservoir of genetic diversity within the genome and exhibits a strong association with economically valuable traits in livestock. The manifestation of aggressive behavior in pigs has detrimental effects on production efficiency, immune competency, and meat quality. Nevertheless, the impact of CNV on the aggressive behavior of pigs remains elusive. In this investigation, we employed an integrated analysis of genome and transcriptome data to investigate the interplay between CNV, gene expression changes, and indicators of aggressive behavior in weaned pigs. Specifically, a subset of pigs comprising the most aggressive pigs (MAP, n = 12) and the least aggressive pigs (LAP, n = 11) was purposefully selected from a herd of 500 weaned pigs following a mixing procedure based on their composite aggressive score (CAS). Subsequently, we thoroughly analyzed copy number variation regions (CNVRs) across the entire genome using next-generation sequencing techniques, ultimately revealing the presence of 6869 CNVRs. Using genome-wide association study (GWAS) analysis and evaluating variance-stabilizing transformation (VST) values, we successfully identified distinct CNVRs that distinguished the MAP and LAP counterparts. Among the prioritized CNVRs, CNVR-4962 (designated as the top-ranked p-value and VST value, No. 1) was located within the Solute Carrier Organic Anion Transporter Family Member 3A1 (SLCO3A1) gene. The results of our analyses indicated a significantly higher (p < 0.05) copy number of SLCO3A1 in the MAP compared to the LAP. Furthermore, this increased copy number exhibited a positive correlation with the CAS of the pigs (p < 0.05). Furthermore, we integrated genomic data with transcriptomic data from the temporal lobe to facilitate the examination of expression quantitative trait loci (eQTL). Importantly, these observations were consistent with the mRNA expression pattern of SLCO3A1 in the temporal lobe of both MAP and LAP (p < 0.05). Consequently, our findings strongly suggest that CNVs affecting SLCO3A1 may influence gene expression through a dosage effect. These results highlight the potential of SLCO3A1 as a candidate gene associated with aggressive traits in pig breeding programs.
Keywords: CNV; SLCO3A1; aggressive behavior; animal welfare; dosage effect.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Genetic Contributions to Aggressive Behaviour in Pigs: A Comprehensive Review.Genes (Basel). 2025 Apr 29;16(5):534. doi: 10.3390/genes16050534. Genes (Basel). 2025. PMID: 40428356 Free PMC article. Review.
-
A composite strategy of genome-wide association study and copy number variation analysis for carcass traits in a Duroc pig population.BMC Genomics. 2022 Aug 13;23(1):590. doi: 10.1186/s12864-022-08804-1. BMC Genomics. 2022. PMID: 35964005 Free PMC article.
-
Genome-wide detection of CNV regions and their potential association with growth and fatness traits in Duroc pigs.BMC Genomics. 2021 May 8;22(1):332. doi: 10.1186/s12864-021-07654-7. BMC Genomics. 2021. PMID: 33964879 Free PMC article.
-
A comprehensive survey of copy number variation in 18 diverse pig populations and identification of candidate copy number variable genes associated with complex traits.BMC Genomics. 2012 Dec 27;13:733. doi: 10.1186/1471-2164-13-733. BMC Genomics. 2012. PMID: 23270433 Free PMC article.
-
Genome wide copy number variations using Porcine 60K SNP Beadchip in Landlly pigs.Anim Biotechnol. 2023 Nov;34(6):1891-1899. doi: 10.1080/10495398.2022.2056047. Epub 2022 Apr 4. Anim Biotechnol. 2023. PMID: 35369845 Review.
Cited by
-
Genetic Contributions to Aggressive Behaviour in Pigs: A Comprehensive Review.Genes (Basel). 2025 Apr 29;16(5):534. doi: 10.3390/genes16050534. Genes (Basel). 2025. PMID: 40428356 Free PMC article. Review.
-
Integrating Deep Learning and Transcriptomics to Assess Livestock Aggression: A Scoping Review.Biology (Basel). 2025 Jun 26;14(7):771. doi: 10.3390/biology14070771. Biology (Basel). 2025. PMID: 40723331 Free PMC article. Review.
References
-
- Turner S.P., Roehe R., D’Eath R.B., Ison S.H., Farish M., Jack M.C., Lundeheim N., Rydhmer L., Lawrence A.B. Genetic validation of postmixing skin injuries in pigs as an indicator of aggressiveness and the relationship with injuries under more stable social conditions. J. Anim. Sci. 2009;87:3076–3082. doi: 10.2527/jas.2008-1558. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

