Super-Resolution Imaging of Neuronal Structures with Structured Illumination Microscopy
- PMID: 37760183
- PMCID: PMC10525192
- DOI: 10.3390/bioengineering10091081
Super-Resolution Imaging of Neuronal Structures with Structured Illumination Microscopy
Abstract
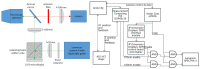
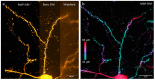
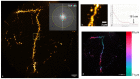
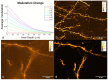
Super-resolution structured illumination microscopy (SR-SIM) is an optical fluorescence microscopy method which is suitable for imaging a wide variety of cells and tissues in biological and biomedical research. Typically, SIM methods use high spatial frequency illumination patterns generated by laser interference. This approach provides high resolution but is limited to thin samples such as cultured cells. Using a different strategy for processing raw data and coarser illumination patterns, we imaged through a 150-micrometer-thick coronal section of a mouse brain expressing GFP in a subset of neurons. The resolution reached 144 nm, an improvement of 1.7-fold beyond conventional widefield imaging.
Keywords: Bayesian methods; brain; fluorescence microscopy; structured illumination; super-resolution.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Update of
-
Super-resolution imaging of neuronal structure with structured illumination microscopy.bioRxiv [Preprint]. 2023 May 26:2023.05.26.542523. doi: 10.1101/2023.05.26.542523. bioRxiv. 2023. Update in: Bioengineering (Basel). 2023 Sep 13;10(9):1081. doi: 10.3390/bioengineering10091081. PMID: 37292949 Free PMC article. Updated. Preprint.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

