TDP-43 Epigenetic Facets and Their Neurodegenerative Implications
- PMID: 37762112
- PMCID: PMC10530927
- DOI: 10.3390/ijms241813807
TDP-43 Epigenetic Facets and Their Neurodegenerative Implications
Abstract
Since its initial involvement in numerous neurodegenerative pathologies in 2006, either as a principal actor or as a cofactor, new pathologies implicating transactive response (TAR) DNA-binding protein 43 (TDP-43) are regularly emerging also beyond the neuronal system. This reflects the fact that TDP-43 functions are particularly complex and broad in a great variety of human cells. In neurodegenerative diseases, this protein is often pathologically delocalized to the cytoplasm, where it irreversibly aggregates and is subjected to various post-translational modifications such as phosphorylation, polyubiquitination, and cleavage. Until a few years ago, the research emphasis has been focused particularly on the impacts of this aggregation and/or on its widely described role in complex RNA splicing, whether related to loss- or gain-of-function mechanisms. Interestingly, recent studies have strengthened the knowledge of TDP-43 activity at the chromatin level and its implication in the regulation of DNA transcription and stability. These discoveries have highlighted new features regarding its own transcriptional regulation and suggested additional mechanistic and disease models for the effects of TPD-43. In this review, we aim to give a comprehensive view of the potential epigenetic (de)regulations driven by (and driving) this multitask DNA/RNA-binding protein.
Keywords: ALS; DNA repair; FTD/FTLD; TARDBP; TDP-43; chromatin; epigenetics; neurodegeneration; retrotransposon; transcriptional regulation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


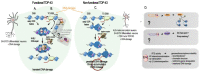
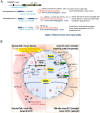



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

