Enriched Aptamer Libraries in Fluorescence-Based Assays for Rikenella microfusus-Specific Gut Microbiome Analyses
- PMID: 37764110
- PMCID: PMC10535755
- DOI: 10.3390/microorganisms11092266
Enriched Aptamer Libraries in Fluorescence-Based Assays for Rikenella microfusus-Specific Gut Microbiome Analyses
Abstract
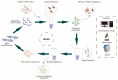
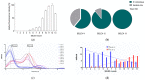
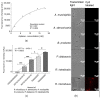
Rikenella microfusus is an essential intestinal probiotic with great potential. The latest research shows that imbalance in the intestinal flora are related to the occurrence of various diseases, such as intestinal diseases, immune diseases, and metabolic diseases. Rikenella may be a target or biomarker for some diseases, providing a new possibility for preventing and treating these diseases by monitoring and optimizing the abundance of Rikenella in the intestine. However, the current monitoring methods have disadvantages, such as long detection times, complicated operations, and high costs, which seriously limit the possibility of clinical application of microbiome-based treatment options. Therefore, the intention of this study was to evolve an enriched aptamer library to be used for specific labeling of R. microfusus, allowing rapid and low-cost detection methods and, ultimately the construction of aptamer-based biosensors. In this study, we used Rikenella as the target bacterium for an in vitro whole Cell-SELEX (Systematic Evolution of Ligands by EXponential Enrichment) to evolve and enrich specific DNA oligonucleotide aptamers. Five other prominent anaerobic gut bacteria were included in this process for counterselection and served as control cells. The aptamer library R.m-R13 was evolved with high specificity and strong affinity (Kd = 9.597 nM after 13 rounds of selection). With this enriched aptamer library, R. microfusus could efficiently be discriminated from the control bacteria in complex mixtures using different analysis techniques, including fluorescence microscopy or fluorometric suspension assays, and even in human stool samples. These preliminary results open new avenues toward the development of aptamer-based microbiome bio-sensing applications for fast and reliable monitoring of R. microfusus.
Keywords: Cell-SELEX; DNA aptamer; Rikenella microfusus; biosensor; in vitro diagnostic.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
A Polyclonal Selex Aptamer Library Directly Allows Specific Labelling of the Human Gut Bacterium Blautia producta without Isolating Individual Aptamers.Molecules. 2022 Sep 3;27(17):5693. doi: 10.3390/molecules27175693. Molecules. 2022. PMID: 36080459 Free PMC article.
-
A Polyclonal Aptamer Library for the Specific Binding of the Gut Bacterium Roseburia intestinalis in Mixtures with Other Gut Microbiome Bacteria and Human Stool Samples.Int J Mol Sci. 2022 Jul 13;23(14):7744. doi: 10.3390/ijms23147744. Int J Mol Sci. 2022. PMID: 35887092 Free PMC article.
-
Polyclonal Aptamers for Specific Fluorescence Labeling and Quantification of the Health Relevant Human Gut Bacterium Parabacteroides distasonis.Microorganisms. 2021 Nov 2;9(11):2284. doi: 10.3390/microorganisms9112284. Microorganisms. 2021. PMID: 34835410 Free PMC article.
-
Advances in Aptamer-Based Biosensors and Cell-Internalizing SELEX Technology for Diagnostic and Therapeutic Application.Biosensors (Basel). 2022 Oct 25;12(11):922. doi: 10.3390/bios12110922. Biosensors (Basel). 2022. PMID: 36354431 Free PMC article. Review.
-
Recent advances on aptamer-based biosensors for detection of pathogenic bacteria.World J Microbiol Biotechnol. 2021 Feb 8;37(3):45. doi: 10.1007/s11274-021-03002-9. World J Microbiol Biotechnol. 2021. PMID: 33554321 Review.
Cited by
-
Effects of Dietary Pretreatment with All-trans Lycopene on Lipopolysaccharide-Induced Jejunal Inflammation: A Multi-Pathway Phenomenon.Foods. 2025 Feb 26;14(5):794. doi: 10.3390/foods14050794. Foods. 2025. PMID: 40077496 Free PMC article.
-
High-Energy Supplemental Feeding Shifts Gut Microbiota Composition and Function in Red Deer (Cervus elaphus).Animals (Basel). 2024 May 10;14(10):1428. doi: 10.3390/ani14101428. Animals (Basel). 2024. PMID: 38791646 Free PMC article.
-
Luteolin Alleviates Ulcerative Colitis in Mice by Modulating Gut Microbiota and Plasma Metabolism.Nutrients. 2025 Jan 7;17(2):203. doi: 10.3390/nu17020203. Nutrients. 2025. PMID: 39861331 Free PMC article.
-
Advancements in the investigation of gut microbiota-based strategies for stroke prevention and treatment.Front Immunol. 2025 Mar 4;16:1533343. doi: 10.3389/fimmu.2025.1533343. eCollection 2025. Front Immunol. 2025. PMID: 40103814 Free PMC article. Review.
-
IMPATIENT-qPCR: monitoring SELEX success during in vitro aptamer evolution.Appl Microbiol Biotechnol. 2024 Apr 4;108(1):284. doi: 10.1007/s00253-024-13085-7. Appl Microbiol Biotechnol. 2024. PMID: 38573322 Free PMC article.
References
-
- CHOJI KANEUCHI T.M. Bacteroides microfusus, a New Species from the Intestines of Calves, Chickens, and Japanese Quails. Int. J. Syst. Evol. Microbiol. 1978;28:478–481. doi: 10.1099/00207713-28-4-478. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

