Halanaerobium polyolivorans sp. nov.-A Novel Halophilic Alkalitolerant Bacterium Capable of Polyol Degradation: Physiological Properties and Genomic Insights
- PMID: 37764169
- PMCID: PMC10536098
- DOI: 10.3390/microorganisms11092325
Halanaerobium polyolivorans sp. nov.-A Novel Halophilic Alkalitolerant Bacterium Capable of Polyol Degradation: Physiological Properties and Genomic Insights
Abstract
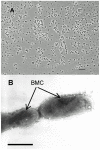
A search for the microorganisms responsible for the anaerobic degradation of osmoprotectants in soda lakes resulted in the isolation of a novel halophilic and alkalitolerant strain, designated Z-7514T. The cells were Gram-stain-negative and non-endospore-forming rods. Optimal growth occurs at 1.6-2.1 M Na+, pH 8.0-8.5, and 31-35 °C. The strain utilized mainly sugars, low molecular polyols, and ethanolamine as well. The G+C content of the genomic DNA of strain Z-7514T was 33.3 mol%. Phylogenetic and phylogenomic analyses revealed that strain Z-7514T belongs to the genus Halanaerobium. On the basis of phenotypic properties and the dDDH and ANI values with close validly published species, it was proposed to evolve strain Z-7514T within the genus Halanaerobium into novel species, for which the name Halanaerobium polyolivorans sp. nov. was proposed. The type strain was Z-7514T (=KCTC 25405T = VKM B-3577T). For species of the genus Halanaerobium, the utilization of ethylene glycol, propylene glycol, and ethanolamine were shown for the first time. The anaerobic degradation of glycols and ethanolamine by strain Z-7514T may represent a novel metabiotic pathway within the alkaliphilic microbial community. Based on a detailed genomic analysis, the main pathways of catabolism of most of the used substrates have been identified.
Keywords: Halanaerobium; alkaliphilic microbial community; ethanolamine; glycerol; polyol degradation.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures



Similar articles
-
Isachenkonia alkalipeptolytica gen. nov. sp. nov., a new anaerobic, alkaliphilic proteolytic bacterium capable of reducing Fe(III) and sulfur.Int J Syst Evol Microbiol. 2020 Aug;70(8):4730-4738. doi: 10.1099/ijsem.0.004341. Epub 2020 Jul 22. Int J Syst Evol Microbiol. 2020. PMID: 32697189
-
Natranaerobaculum magadiense gen. nov., sp. nov., an anaerobic, alkalithermophilic bacterium from soda lake sediment.Int J Syst Evol Microbiol. 2013 Dec;63(Pt 12):4456-4461. doi: 10.1099/ijs.0.054536-0. Epub 2013 Jul 16. Int J Syst Evol Microbiol. 2013. PMID: 23859946
-
Fuchsiella alkaliacetigena gen. nov., sp. nov., an alkaliphilic, lithoautotrophic homoacetogen from a soda lake.Int J Syst Evol Microbiol. 2012 Jul;62(Pt 7):1666-1673. doi: 10.1099/ijs.0.034363-0. Epub 2011 Sep 9. Int J Syst Evol Microbiol. 2012. PMID: 21908678
-
Halanaerobium sehlinense sp. nov., an extremely halophilic, fermentative, strictly anaerobic bacterium from sediments of the hypersaline lake Sehline Sebkha.Int J Syst Evol Microbiol. 2013 Jun;63(Pt 6):2069-2074. doi: 10.1099/ijs.0.040139-0. Epub 2012 Oct 12. Int J Syst Evol Microbiol. 2013. PMID: 23064350
-
Desulfonatronum cooperativum sp. nov., a novel hydrogenotrophic, alkaliphilic, sulfate-reducing bacterium, from a syntrophic culture growing on acetate.Int J Syst Evol Microbiol. 2005 May;55(Pt 3):1001-1006. doi: 10.1099/ijs.0.63490-0. Int J Syst Evol Microbiol. 2005. PMID: 15879225
Cited by
-
Diversity, Taxonomic Novelty, and Encoded Functions of Salar de Ascotán Microbiota, as Revealed by Metagenome-Assembled Genomes.Microorganisms. 2023 Nov 20;11(11):2819. doi: 10.3390/microorganisms11112819. Microorganisms. 2023. PMID: 38004830 Free PMC article.
References
-
- Grant W.D., Jones B.E. Bacteria, archaea and viruses of soda lakes. In: Schagerl M., editor. Soda Lakes of East Africa. Springer International Publishing; Cham, Switzerland: 2016. pp. 97–147. - DOI
-
- Zhilina T.N., Zavarzin G.A. Alkaliphilic anaerobic community at pH 10. Curr. Microbiol. 1994;28:109–112. doi: 10.1007/BF01575757. - DOI
-
- Boltyanskaya Y., Kevbrin V.V. Laboratory simulation of “Proteolytic bacterium—Cyanobacterium” interaction in alkaliphilic microbial community. Paleontol. J. 2018;52:1179–1185. doi: 10.1134/S0031030118100064. - DOI
LinkOut - more resources
Full Text Sources

