Identification and Expression of the MADS-box Gene Family in Different Versions of the Ginkgo biloba Genome
- PMID: 37765498
- PMCID: PMC10535167
- DOI: 10.3390/plants12183334
Identification and Expression of the MADS-box Gene Family in Different Versions of the Ginkgo biloba Genome
Abstract
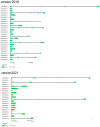
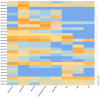
MADS-box transcription factors play important roles in many organisms. These transcription factors are involved in processes such as the formation of the flower organ structure and the seed development of plants. Ginkgo biloba has two genome versions (version 2019 and version 2021), and there is no analysis or comparison of the MADS-box gene family in these two genomes. In this study, 26 and 20 MADS-box genes were identified from the two genomes of Ginkgo, of which 12 pairs of genes reached more than 80% similarity. According to our phylogenetic analysis results, we divided these genes into type I (Mα and Mγ subfamilies) and type II (MIKC and Mδ subfamilies) members. We found that both sets of genomes lacked the Mβ gene, while the MIKC gene was the most numerous. Further analysis of the gene structure showed that the MIKC genes in the two genomes had extralong introns (≥20 kb); these introns had different splicing patterns, and their expression might be more abundant. The gene expression analysis proved that GbMADS genes were expressed to varying degrees in eight Ginkgo biological tissues. Type II GbMADS genes not only were found to be related to female flower bud differentiation and development but also are important in seed development. Therefore, MADS-box genes may play important roles in the development of Ginkgo reproductive organs, which may suggest a genetic role in sexual differentiation. This study further contributes to the research on MADS-box genes and provides new insights into sex determination in Ginkgo.
Keywords: Ginkgo genome; MADS-box; expression; gene structure.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Genome-wide identification and classification of MIKC-type MADS-box genes in Streptophyte lineages and expression analyses to reveal their role in seed germination of orchid.BMC Plant Biol. 2019 May 28;19(1):223. doi: 10.1186/s12870-019-1836-5. BMC Plant Biol. 2019. PMID: 31138149 Free PMC article.
-
Genome-wide identification, characterisation and expression analysis of the MADS-box gene family in Prunus mume.Mol Genet Genomics. 2014 Oct;289(5):903-20. doi: 10.1007/s00438-014-0863-z. Epub 2014 May 25. Mol Genet Genomics. 2014. PMID: 24859011
-
Comparative Analysis of the MADS-Box Genes Revealed Their Potential Functions for Flower and Fruit Development in Longan (Dimocarpus longan).Front Plant Sci. 2022 Jan 27;12:813798. doi: 10.3389/fpls.2021.813798. eCollection 2021. Front Plant Sci. 2022. PMID: 35154209 Free PMC article.
-
Genome-wide Analysis of the MADS-Box Gene Family in Watermelon.Comput Biol Chem. 2019 Jun;80:341-350. doi: 10.1016/j.compbiolchem.2019.04.013. Epub 2019 May 2. Comput Biol Chem. 2019. PMID: 31082717 Review.
-
The major clades of MADS-box genes and their role in the development and evolution of flowering plants.Mol Phylogenet Evol. 2003 Dec;29(3):464-89. doi: 10.1016/s1055-7903(03)00207-0. Mol Phylogenet Evol. 2003. PMID: 14615187 Review.
References
-
- Parenicova L., de Folter S., Kieffer M., Horner D.S., Favalli C., Busscher J., Cook H.E., Ingram R.M., Kater M.M., Davies B., et al. Molecular and phylogenetic analyses of the complete MADS-box transcription factor family in Arabidopsis: New openings to the MADS world. Plant Cell. 2003;15:1538–1551. doi: 10.1105/tpc.011544. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

