Curved adhesions mediate cell attachment to soft matrix fibres in three dimensions
- PMID: 37770566
- PMCID: PMC10567576
- DOI: 10.1038/s41556-023-01238-1
Curved adhesions mediate cell attachment to soft matrix fibres in three dimensions
Abstract
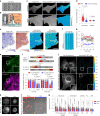
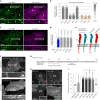
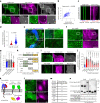
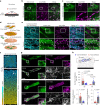
Integrin-mediated focal adhesions are the primary architectures that transmit forces between the extracellular matrix (ECM) and the actin cytoskeleton. Although focal adhesions are abundant on rigid and flat substrates that support high mechanical tensions, they are sparse in soft three-dimensional (3D) environments. Here we report curvature-dependent integrin-mediated adhesions called curved adhesions. Their formation is regulated by the membrane curvatures imposed by the topography of ECM protein fibres. Curved adhesions are mediated by integrin ɑvβ5 and are molecularly distinct from focal adhesions and clathrin lattices. The molecular mechanism involves a previously unknown interaction between integrin β5 and a curvature-sensing protein, FCHo2. We find that curved adhesions are prevalent in physiological conditions, and disruption of curved adhesions inhibits the migration of some cancer cell lines in 3D fibre matrices. These findings provide a mechanism for cell anchorage to natural protein fibres and suggest that curved adhesions may serve as a potential therapeutic target.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





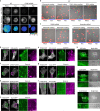

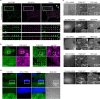
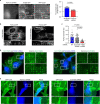


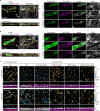
Update of
-
Curved adhesions mediate cell attachment to soft matrix fibres in 3D.bioRxiv [Preprint]. 2023 Mar 19:2023.03.16.532975. doi: 10.1101/2023.03.16.532975. bioRxiv. 2023. Update in: Nat Cell Biol. 2023 Oct;25(10):1453-1464. doi: 10.1038/s41556-023-01238-1. PMID: 36993504 Free PMC article. Updated. Preprint.
References
-
- Klapholz B, Brown NH. Talin—the master of integrin adhesions. J. Cell Sci. 2017;130:2435–2446. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

