Chromatin accessibility profiling of targeted cell populations with laser capture microdissection coupled to ATAC-seq
- PMID: 37776856
- PMCID: PMC10626193
- DOI: 10.1016/j.crmeth.2023.100598
Chromatin accessibility profiling of targeted cell populations with laser capture microdissection coupled to ATAC-seq
Abstract
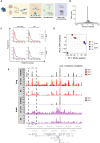
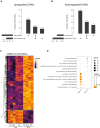
Spatially resolved omics technologies reveal context-dependent cellular regulatory networks in tissues of interest. Beyond transcriptome analysis, information on epigenetic traits and chromatin accessibility can provide further insights on gene regulation in health and disease. Nevertheless, compared to the enormous advancements in spatial transcriptomics technologies, the field of spatial epigenomics is much younger and still underexplored. In this study, we report laser capture microdissection coupled to ATAC-seq (LCM-ATAC-seq) applied to fresh frozen samples for the spatial characterization of chromatin accessibility. We first demonstrate the efficient use of LCM coupled to in situ tagmentation and evaluate its performance as a function of cell number, microdissected areas, and tissue type. Further, we demonstrate its use for the targeted chromatin accessibility analysis of discrete contiguous or scattered cell populations in tissues via single-nuclei capture based on immunostaining for specific cellular markers.
Keywords: ATAC-seq; CP: Molecular biology; chromatin accessibility; epigenomics; laser capture microdissection; spatial omics.
Copyright © 2023 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures



References
-
- Madissoon E., Oliver A.J., Kleshchevnikov V., Wilbrey-Clark A., Polanski K., Richoz N., Ribeiro Orsi A., Mamanova L., Bolt L., Elmentaite R., et al. A spatially resolved atlas of the human lung characterizes a gland-associated immune niche. Nat. Genet. 2023;55:66–77. doi: 10.1038/s41588-022-01243-4. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

