This is a preprint.
Systematic Perturbation of Thousands of Retroviral LTRs in Mouse Embryos
- PMID: 37781606
- PMCID: PMC10541133
- DOI: 10.1101/2023.09.19.558531
Systematic Perturbation of Thousands of Retroviral LTRs in Mouse Embryos
Update in
-
Systematic evaluation of retroviral LTRs as cis-regulatory elements in mouse embryos.Cell Rep. 2024 Mar 26;43(3):113775. doi: 10.1016/j.celrep.2024.113775. Epub 2024 Feb 20. Cell Rep. 2024. PMID: 38381606 Free PMC article.
Abstract
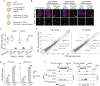
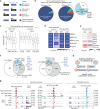
In mammals, many retrotransposons are de-repressed during zygotic genome activation (ZGA). However, their functions in early development remain elusive largely due to the challenge to simultaneously manipulate thousands of retrotransposon insertions in embryos. Here, we employed epigenome editing to perturb the long terminal repeat (LTR) MT2_Mm, a well-known ZGA and totipotency marker that exists in ~2667 insertions throughout the mouse genome. CRISPRi robustly repressed 2485 (~93%) MT2_Mm insertions and 1090 (~55%) insertions of the closely related MT2C_Mm in 2-cell embryos. Remarkably, such perturbation caused down-regulation of hundreds of ZGA genes at the 2-cell stage and embryonic arrest mostly at the morula stage. Mechanistically, MT2_Mm/MT2C_Mm primarily served as alternative ZGA promoters activated by OBOX proteins. Thus, through unprecedented large-scale epigenome editing, we addressed to what extent MT2_Mm/MT2C_Mm regulates ZGA and preimplantation development. Our approach could be adapted to systematically perturb retrotransposons in other mammalian embryos as it doesn't require transgenic animals.
Keywords: MERVL; MT2_Mm; endogenous retroviruses; epigenome editing; long terminal repeats; preimplantation embryos; zygotic genome activation.
Conflict of interest statement
9.Author information The authors declare no competing financial interests.
Figures


References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
