Molecular analysis of the type III interferon complex and its applications in protein engineering
- PMID: 37794680
- PMCID: PMC10645568
- DOI: 10.1016/j.bpj.2023.09.021
Molecular analysis of the type III interferon complex and its applications in protein engineering
Abstract
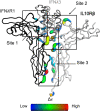
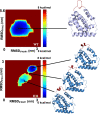
Type III interferons (IFNλs) are cytokines with critical roles in the immune system and are attractive therapeutic candidates due to their tissue-specific activity. Despite entering several clinical trials, results have demonstrated limited efficacy and potency, partially attributed to low-affinity protein-protein interactions (PPIs) responsible for receptor complex formation. Subsequently, structural studies of the native IFNλ signaling complexes remain inaccessible. While protein engineering can overcome affinity limitations, tools to investigate low-affinity systems like these remain limited. To provide insights into previous efforts to strengthen the PPIs within this complex, we perform a molecular analysis of the extracellular ternary complexes of IFNλ3 using both computational and experimental approaches. We first use molecular simulations and modeling to quantify differences in PPIs and residue strain fluctuations, generate detailed free energy landscapes, and reveal structural differences between an engineered, high-affinity complex, and a model of the wild-type, low-affinity complex. This analysis illuminates distinct behaviors of these ligands, yielding mechanistic insights into IFNλ complex formation. We then apply these computational techniques in protein engineering and design by utilizing simulation data to identify hotspots of interaction to rationally engineer the native cytokine-receptor complex for increased stability. These simulations are then validated by experimental techniques, showing that a single mutation at a computationally predicted site of interaction between the two receptors increases PPIs and improves complex formation for all IFNλs. This study highlights the power of molecular dynamics simulations for protein engineering and design as applied to the IFNλ family but also presents a potential tool for analysis and engineering of other systems with low-affinity PPIs.
Copyright © 2023 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests at this time.
Figures


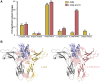

References
-
- Packer M.S., Liu D.R. Methods for the directed evolution of proteins. Nat. Rev. Genet. 2015;16:379–394. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

