Unlatching of the stem domains in the Staphylococcus aureus pore-forming leukocidin LukAB influences toxin oligomerization
- PMID: 37802313
- PMCID: PMC10665946
- DOI: 10.1016/j.jbc.2023.105321
Unlatching of the stem domains in the Staphylococcus aureus pore-forming leukocidin LukAB influences toxin oligomerization
Abstract
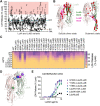
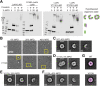
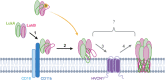
Staphylococcus aureus (S. aureus) is a serious global pathogen that causes a diverse range of invasive diseases. S. aureus utilizes a family of pore-forming toxins, known as bi-component leukocidins, to evade the host immune response and promote infection. Among these is LukAB (leukocidin A/leukocidin B), a toxin that assembles into an octameric β-barrel pore in the target cell membrane, resulting in host cell death. The established cellular receptor for LukAB is CD11b of the Mac-1 complex. Here, we show that hydrogen voltage-gated channel 1 is also required for the cytotoxicity of all major LukAB variants. We demonstrate that while each receptor is sufficient to recruit LukAB to the plasma membrane, both receptors are required for maximal lytic activity. Why LukAB requires two receptors, and how each of these receptors contributes to pore-formation remains unknown. To begin to resolve this, we performed an alanine scanning mutagenesis screen to identify mutations that allow LukAB to maintain cytotoxicity without CD11b. We discovered 30 mutations primarily localized in the stem domains of LukA and LukB that enable LukAB to exhibit full cytotoxicity in the absence of CD11b. Using crosslinking, electron microscopy, and hydroxyl radical protein footprinting, we show these mutations increase the solvent accessibility of the stem domain, priming LukAB for oligomerization. Together, our data support a model in which CD11b binding unlatches the membrane penetrating stem domains of LukAB, and this change in flexibility promotes toxin oligomerization.
Keywords: Staphylococcus aureus (S. aureus); bacterial toxin; high-throughput screening; oligomerization; receptor.
Copyright © 2023 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest V. J. T. is an inventor on patents and patent applications filed by New York University, which are currently under commercial license to Janssen Biotech Inc. Janssen Biotech provides research funding and other payments associated with the licensing agreement. All other authors declare no conflict of interest.
Figures



Similar articles
-
Identification of a crucial residue required for Staphylococcus aureus LukAB cytotoxicity and receptor recognition.Infect Immun. 2014 Mar;82(3):1268-76. doi: 10.1128/IAI.01444-13. Epub 2013 Dec 30. Infect Immun. 2014. PMID: 24379286 Free PMC article.
-
Staphylococcus aureus LukAB cytotoxin kills human neutrophils by targeting the CD11b subunit of the integrin Mac-1.Proc Natl Acad Sci U S A. 2013 Jun 25;110(26):10794-9. doi: 10.1073/pnas.1305121110. Epub 2013 Jun 10. Proc Natl Acad Sci U S A. 2013. PMID: 23754403 Free PMC article.
-
Staphylococcus aureus Biofilms Induce Macrophage Dysfunction Through Leukocidin AB and Alpha-Toxin.mBio. 2015 Aug 25;6(4):e01021-15. doi: 10.1128/mBio.01021-15. mBio. 2015. PMID: 26307164 Free PMC article.
-
Structure and Function of the Two-Component Cytotoxins of Staphylococcus aureus - Learnings for Designing Novel Therapeutics.Adv Exp Med Biol. 2017;966:15-35. doi: 10.1007/5584_2016_200. Adv Exp Med Biol. 2017. PMID: 28455832 Review.
-
Staphylococcus aureus hemolysins, bi-component leukocidins, and cytolytic peptides: a redundant arsenal of membrane-damaging virulence factors?Front Cell Infect Microbiol. 2012 Feb 16;2:12. doi: 10.3389/fcimb.2012.00012. eCollection 2012. Front Cell Infect Microbiol. 2012. PMID: 22919604 Free PMC article. Review.
References
-
- King M.D., Humphrey B.J., Wang Y.F., Kourbatova E.V., Ray S.M., Blumberg H.M. Emergence of community-acquired methicillin-resistant Staphylococcus aureus USA 300 clone as the predominant cause of skin and soft-tissue infections. Ann. Intern. Med. 2006;144:309–317. - PubMed
-
- Witte W., Kresken M., Braulke C., Cuny C. Increasing incidence and widespread dissemination of methicillin-resistant Staphylococcus aureus (MRSA) in hospitals in central Europe, with special reference to German hospitals. Clin. Microbiol. Infect. 1997;3:414–422. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

