Acute myeloid leukemia: from NGS, through scRNA-seq, to CAR-T. dissect cancer heterogeneity and tailor the treatment
- PMID: 37803464
- PMCID: PMC10557350
- DOI: 10.1186/s13046-023-02841-8
Acute myeloid leukemia: from NGS, through scRNA-seq, to CAR-T. dissect cancer heterogeneity and tailor the treatment
Abstract
Acute myeloid leukemia (AML) is a malignant blood cancer with marked cellular heterogeneity due to altered maturation and differentiation of myeloid blasts, the possible causes of which are transcriptional or epigenetic alterations, impaired apoptosis, and excessive cell proliferation. This neoplasm has a high rate of resistance to anticancer therapies and thus a high risk of relapse and mortality because of both the biological diversity of the patient and intratumoral heterogeneity due to the acquisition of new somatic changes. For more than 40 years, the old gold standard "one size fits all" treatment approach included intensive chemotherapy treatment with anthracyclines and cytarabine.The manuscript first traces the evolution of the understanding of the pathology from the 1970s to the present. The enormous strides made in its categorization prove to be crucial for risk stratification, enabling an increasingly personalized diagnosis and treatment approach.Subsequently, we highlight how, over the past 15 years, technological advances enabling single cell RNA sequencing and T-cell modification based on the genomic tools are affecting the classification and treatment of AML. At the dawn of the new millennium, the advent of high-throughput next-generation sequencing technologies has enabled the profiling of patients evidencing different facets of the same disease, stratifying risk, and identifying new possible therapeutic targets that have subsequently been validated. Currently, the possibility of investigating tumor heterogeneity at the single cell level, profiling the tumor at the time of diagnosis or after treatments exist. This would allow the identification of underrepresented cellular subclones or clones resistant to therapeutic approaches and thus responsible for post-treatment relapse that would otherwise be difficult to detect with bulk investigations on the tumor biopsy. Single-cell investigation will then allow even greater personalization of therapy to the genetic and transcriptional profile of the tumor, saving valuable time and dangerous side effects. The era of personalized medicine will take a huge step forward through the disclosure of each individual piece of the complex puzzle that is cancer pathology, to implement a "tailored" therapeutic approach based also on engineered CAR-T cells.
Keywords: Acute myeloid leukaemia (AML); Intratumoral heterogeneity; Next generation sequencing (NGS); Personalized medicine; Single cell sequencing (scRNA-seq); Therapeutic targets.
© 2023. Italian National Cancer Institute ‘Regina Elena’.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


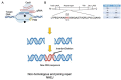
References
-
- D. P. Hansemann. (Virchow’s Arch., 1890), vol. 119, pp. 299–326.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

