This is a preprint.
Bacterial cell wall biosynthesis is controlled by growth rate dependent modulation of turgor pressure in E. coli
- PMID: 37808635
- PMCID: PMC10557573
- DOI: 10.1101/2023.08.31.555748
Bacterial cell wall biosynthesis is controlled by growth rate dependent modulation of turgor pressure in E. coli
Abstract
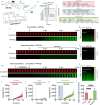
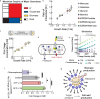
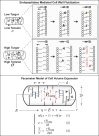
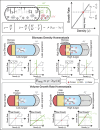
The cell wall is an essential cellular component of bacteria and the target of many antibiotics. However, how bacteria regulate the rate of cell wall biosynthesis as growth rates change remains unresolved. In E. coli, cell wall growth was thought to proceed independently from turgor pressure1, the osmotic pressure that the cytoplasm exerts on the cell wall. Here, we uncover a striking increase of turgor pressure with growth rate. Modulating turgor pressure and measuring cell wall biosynthesis, we find that turgor pressure is directly controls the rate of cell wall biosynthesis. The picture that emerges is that turgor pressure is largely generated by counterions of negatively charged cellular biomass. The increase in turgor pressure with growth rates results from more ribosomes and therefore higher concentrations of negatively charged ribosomal RNA. Elegantly, the coupling between biomass composition, turgor pressure and cell wall biosynthesis simultaneously explains how bacteria achieve homeostasis of cytoplasmic crowding and how they regulate the rate of cell wall biosynthesis across growth rates.
Figures



Similar articles
-
Homeostasis of cytoplasmic crowding by cell wall fluidization and ribosomal counterions.Res Sq [Preprint]. 2024 Apr 19:rs.3.rs-4138690. doi: 10.21203/rs.3.rs-4138690/v1. Res Sq. 2024. PMID: 38699329 Free PMC article. Preprint.
-
Comparison of Two Modern Survival Prediction Tools, SORG-MLA and METSSS, in Patients With Symptomatic Long-bone Metastases Who Underwent Local Treatment With Surgery Followed by Radiotherapy and With Radiotherapy Alone.Clin Orthop Relat Res. 2024 Dec 1;482(12):2193-2208. doi: 10.1097/CORR.0000000000003185. Epub 2024 Jul 23. Clin Orthop Relat Res. 2024. PMID: 39051924
-
Short-Term Memory Impairment.2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31424720 Free Books & Documents.
-
Automated devices for identifying peripheral arterial disease in people with leg ulceration: an evidence synthesis and cost-effectiveness analysis.Health Technol Assess. 2024 Aug;28(37):1-158. doi: 10.3310/TWCG3912. Health Technol Assess. 2024. PMID: 39186036 Free PMC article.
-
Stem cell transplantation for induction of remission in medically refractory Crohn's disease.Cochrane Database Syst Rev. 2022 May 13;5(5):CD013070. doi: 10.1002/14651858.CD013070.pub2. Cochrane Database Syst Rev. 2022. PMID: 35556242 Free PMC article.
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
