This is a preprint.
The Physical Biology of Nucleolus Disassembly
- PMID: 37808669
- PMCID: PMC10557732
- DOI: 10.1101/2023.09.27.559731
The Physical Biology of Nucleolus Disassembly
Abstract
During cell division, precise and regulated distribution of cellular material between daughter cells is a critical step and is governed by complex biochemical and biophysical mechanisms. To achieve this, membraneless organelles and condensates often require complete disassembly during mitosis. The biophysical principles governing the disassembly of condensates remain poorly understood. Here, we used a physical biology approach to study how physical and material properties of the nucleolus, a prominent nuclear membraneless organelle in eukaryotic cells, change during mitosis and across different scales. We found that nucleolus disassembly proceeds continuously through two distinct phases with a slow and reversible preparatory phase followed by a rapid irreversible phase that was concurrent with the nuclear envelope breakdown. We measured microscopic properties of nucleolar material including effective diffusion rates and binding affinities as well as key macroscopic properties of surface tension and bending rigidity. By incorporating these measurements into the framework of critical phenomena, we found evidence that near mitosis surface tension displays a power-law behavior as a function of biochemically modulated interaction strength. This two-step disassembly mechanism, which maintains structural and functional stability of nucleolus while allowing for its rapid and efficient disassembly in response to cell cycle cues, may be a universal design principle for the disassembly of other biomolecular condensates.
Figures



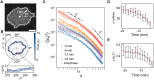

References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
