Tracking gut microbiome and bloodstream infection in critically ill adults
- PMID: 37816004
- PMCID: PMC10564172
- DOI: 10.1371/journal.pone.0289923
Tracking gut microbiome and bloodstream infection in critically ill adults
Abstract
Background: The gut microbiome is believed to contribute to bloodstream infection (BSI) via translocation of dominant gut bacteria in vulnerable patient populations. However, conclusively linking gut and blood organisms requires stringent approaches to establish strain-level identity.
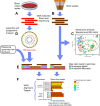
Methods: We enrolled a convenience cohort of critically ill patients and investigated 86 bloodstream infection episodes that occurred in 57 patients. Shotgun metagenomic sequencing was used to define constituents of their gut microbiomes, and whole genome sequencing and assembly was done on 23 unique bloodstream isolates that were available from 21 patients. Whole genome sequences were downloaded from public databases and used to establish sequence-identity distribution and define thresholds for unrelated genomes of BSI species. Gut microbiome reads were then aligned to whole genome sequences of the cognate bloodstream isolate and unrelated database isolates to assess identity.
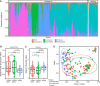
Results: Gut microbiome constituents matching the bloodstream infection species were present in half of BSI episodes, and represented >30% relative abundance of gut sequences in 10% of episodes. Among the 23 unique bloodstream organisms that were available for whole genome sequencing, 14 were present in gut at the species level. Sequence alignment applying defined thresholds for identity revealed that 6 met criteria for identical strains in blood and gut, but 8 did not. Sequence identity between BSI isolates and gut microbiome reads was more likely when the species was present at higher relative abundance in gut.
Conclusion: In assessing potential gut source for BSI, stringent sequence-based approaches are essential to determine if organisms responsible for BSI are identical to those in gut: of 14 evaluable patients in which the same species was present in both sites, they were identical in 6/14, but were non-identical in 8/14 and thus inconsistent with gut source. This report demonstrates application of sequencing as a key tool to investigate infection tracking within patients.
Copyright: © 2023 Gu et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

References
-
- Courjon J, Demonchy E, Degand N, Risso K, Ruimy R, Roger PM. Patients with community-acquired bacteremia of unknown origin: clinical characteristics and usefulness of microbiological results for therapeutic issues: a single-center cohort study. Annals of clinical microbiology and antimicrobials. 2017;16(1):40. Epub 2017/05/21. doi: 10.1186/s12941-017-0214-0 ; PubMed Central PMCID: PMC5438554. - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

