SARS-CoV-2 genomic surveillance in wastewater as a model for monitoring evolution of endemic viruses
- PMID: 37816740
- PMCID: PMC10564906
- DOI: 10.1038/s41467-023-41369-5
SARS-CoV-2 genomic surveillance in wastewater as a model for monitoring evolution of endemic viruses
Abstract
As global SARS-CoV-2 burden and testing frequency have decreased, wastewater surveillance has emerged as a key tool to support clinical surveillance efforts. The aims of this study were to identify and characterize SARS-CoV-2 variants in wastewater samples collected from urban centers across South Africa. Here we show that wastewater sequencing analyses are temporally concordant with clinical genomic surveillance and reveal the presence of multiple lineages not detected by clinical surveillance. We show that wastewater genomics can support SARS-CoV-2 epidemiological investigations by reliably recovering the prevalence of local circulating variants, even when clinical samples are not available. Further, we find that analysis of mutations observed in wastewater can provide a signal of upcoming lineage transitions. Our study demonstrates the utility of wastewater genomics to monitor evolution and spread of endemic viruses.
© 2023. Springer Nature Limited.
Conflict of interest statement
All authors declare no competing interests.
Figures
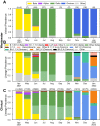
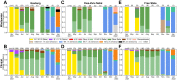
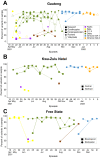

References
-
- WHO. Environmental surveillance for SARS-COV-2 to complement public health surveillance – Interim Guidance. https://www.who.int/publications/i/item/WHO-HEP-ECH-WSH-2022.1 (2022).
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

