Dissection of key factors correlating with H5N1 avian influenza virus driven inflammatory lung injury of chicken identified by single-cell analysis
- PMID: 37819993
- PMCID: PMC10593216
- DOI: 10.1371/journal.ppat.1011685
Dissection of key factors correlating with H5N1 avian influenza virus driven inflammatory lung injury of chicken identified by single-cell analysis
Abstract
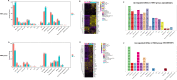
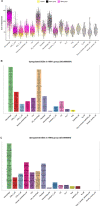
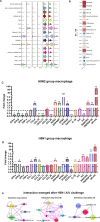
Chicken lung is an important target organ of avian influenza virus (AIV) infection, and different pathogenic virus strains lead to opposite prognosis. Using a single-cell RNA sequencing (scRNA-seq) assay, we systematically and sequentially analyzed the transcriptome of 16 cell types (19 clusters) in the lung tissue of chickens infected with H5N1 highly pathogenic avian influenza virus (HPAIV) and H9N2 low pathogenic avian influenza virus (LPAIV), respectively. Notably, we developed a valuable catalog of marker genes for these cell types. Compared to H9N2 AIV infection, H5N1 AIV infection induced extensive virus replication and the immune reaction across most cell types simultaneously. More importantly, we propose that infiltrating inflammatory macrophages (clusters 0, 1, and 14) with massive viral replication, pro-inflammatory cytokines (IFN-β, IL1β, IL6 and IL8), and emerging interaction of various cell populations through CCL4, CCL19 and CXCL13, potentially contributed to the H5N1 AIV driven inflammatory lung injury. Our data revealed complex but distinct immune response landscapes in the lung tissue of chickens after H5N1 and H9N2 AIV infection, and deciphered the potential mechanisms underlying AIV-driven inflammatory reactions in chicken. Furthermore, this article provides a rich database for the molecular basis of different cell-type responses to AIV infection.
Copyright: © 2023 Dai et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


References
-
- Dai M, Li S, Keyi S, Sun H, Zhao L, Deshui Y, et al.. Comparative analysis of key immune protection factors in H9N2 avian influenza viruses infected and immunized specific pathogen-free chicken. Poultry science. 2021;100(1):39–46. Epub 2020/12/29. doi: 10.1016/j.psj.2020.09.080 ; PubMed Central PMCID: PMC7772655. - DOI - PMC - PubMed
-
- Chen X, Wang W, Wang Y, Lai S, Yang J, Cowling BJ, et al.. Serological evidence of human infections with highly pathogenic avian influenza A(H5N1) virus: a systematic review and meta-analysis. BMC medicine. 2020;18(1):377. doi: 10.1186/s12916-020-01836-y ; PubMed Central PMCID: PMC7709391. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical

