Interindividual variation in human cortical cell type abundance and expression
- PMID: 37824649
- PMCID: PMC11702338
- DOI: 10.1126/science.adf2359
Interindividual variation in human cortical cell type abundance and expression
Abstract
Single-cell transcriptomic studies have identified a conserved set of neocortical cell types from small postmortem cohorts. We extended these efforts by assessing cell type variation across 75 adult individuals undergoing epilepsy and tumor surgeries. Nearly all nuclei map to one of 125 robust cell types identified in the middle temporal gyrus. However, we found interindividual variance in abundances and gene expression signatures, particularly in deep-layer glutamatergic neurons and microglia. A minority of donor variance is explainable by age, sex, ancestry, disease state, and cell state. Genomic variation was associated with expression of 150 to 250 genes for most cell types. This characterization of cellular variation provides a baseline for cell typing in health and disease.
Figures

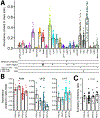



References
-
- Evans AC et al., Anatomical mapping of functional activation in stereotactic coordinate space. Neuroimage. 1, 43–53 (1992). - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

