New Insights into Radio-Resistance Mechanism Revealed by (Phospho)Proteome Analysis of Deinococcus Radiodurans after Heavy Ion Irradiation
- PMID: 37834265
- PMCID: PMC10572868
- DOI: 10.3390/ijms241914817
New Insights into Radio-Resistance Mechanism Revealed by (Phospho)Proteome Analysis of Deinococcus Radiodurans after Heavy Ion Irradiation
Abstract
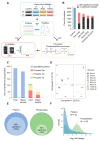
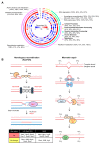
Deinococcus radiodurans (D. radiodurans) can tolerate various extreme environments including radiation. Protein phosphorylation plays an important role in radiation resistance mechanisms; however, there is currently a lack of systematic research on this topic in D. radiodurans. Based on label-free (phospho)proteomics, we explored the dynamic changes of D. radiodurans under various doses of heavy ion irradiation and at different time points. In total, 2359 proteins and 1110 high-confidence phosphosites were identified, of which 66% and 23% showed significant changes, respectively, with the majority being upregulated. The upregulated proteins at different states (different doses or time points) were distinct, indicating that the radio-resistance mechanism is dose- and stage-dependent. The protein phosphorylation level has a much higher upregulation than protein abundance, suggesting phosphorylation is more sensitive to irradiation. There were four distinct dynamic changing patterns of phosphorylation, most of which were inconsistent with protein levels. Further analysis revealed that pathways related to RNA metabolism and antioxidation were activated after irradiation, indicating their importance in radiation response. We also screened some key hub phosphoproteins and radiation-responsive kinases for further study. Overall, this study provides a landscape of the radiation-induced dynamic change of protein expression and phosphorylation, which provides a basis for subsequent functional and applied studies.
Keywords: Deinococcus radiodurans; dynamic change; kinase; phosphoproteome; proteome; radiation.
Conflict of interest statement
The authors declare that they have no competing interest.
Figures




Similar articles
-
iTRAQ-based proteomic analysis of Deinococcus radiodurans in response to 12C6+ heavy ion irradiation.BMC Microbiol. 2022 Nov 4;22(1):264. doi: 10.1186/s12866-022-02676-x. BMC Microbiol. 2022. PMID: 36333788 Free PMC article.
-
Quantitative redox proteomics links thioredoxin to heavy ion resistance in Deinococcus radiodurans.Free Radic Biol Med. 2025 Mar 1;229:225-236. doi: 10.1016/j.freeradbiomed.2024.12.042. Epub 2024 Dec 20. Free Radic Biol Med. 2025. PMID: 39710107
-
Phosphorylation of FtsZ and FtsA by a DNA Damage-Responsive Ser/Thr Protein Kinase Affects Their Functional Interactions in Deinococcus radiodurans.mSphere. 2018 Jul 18;3(4):e00325-18. doi: 10.1128/mSphere.00325-18. mSphere. 2018. PMID: 30021877 Free PMC article.
-
Gene regulation for the extreme resistance to ionizing radiation of Deinococcus radiodurans.Gene. 2019 Oct 5;715:144008. doi: 10.1016/j.gene.2019.144008. Epub 2019 Jul 27. Gene. 2019. PMID: 31362038 Review.
-
Oxidative stress resistance in Deinococcus radiodurans.Microbiol Mol Biol Rev. 2011 Mar;75(1):133-91. doi: 10.1128/MMBR.00015-10. Microbiol Mol Biol Rev. 2011. PMID: 21372322 Free PMC article. Review.
Cited by
-
The Mechanism Underlying the Hypoglycemic Effect of Epimedin C on Mice with Type 2 Diabetes Mellitus Based on Proteomic Analysis.Nutrients. 2023 Dec 21;16(1):25. doi: 10.3390/nu16010025. Nutrients. 2023. PMID: 38201855 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

