The Initial Hepatitis B Virus-Hepatocyte Genomic Integrations and Their Role in Hepatocellular Oncogenesis
- PMID: 37834296
- PMCID: PMC10573506
- DOI: 10.3390/ijms241914849
The Initial Hepatitis B Virus-Hepatocyte Genomic Integrations and Their Role in Hepatocellular Oncogenesis
Abstract
Hepatitis B virus (HBV) remains a dominant cause of hepatocellular carcinoma (HCC). Recently, it was shown that HBV and woodchuck hepatitis virus (WHV) integrate into the hepatocyte genome minutes after invasion. Retrotransposons and transposable sequences were frequent sites of the initial insertions, suggesting a mechanism for spontaneous HBV DNA dispersal throughout the hepatocyte genome. Several somatic genes were also identified as early insertional targets in infected hepatocytes and woodchuck livers. Head-to-tail joints (HTJs) dominated amongst fusions, indicating their creation by non-homologous end-joining (NHEJ). Their formation coincided with the robust oxidative damage of hepatocyte DNA. This was associated with the activation of poly(ADP-ribose) polymerase 1 (PARP1)-mediated dsDNA repair, as reflected by the augmented transcription of PARP1 and XRCC1; the PARP1 binding partner OGG1, a responder to oxidative DNA damage; and increased activity of NAD+, a marker of PARP1 activation, and HO1, an indicator of cell oxidative stress. The engagement of the PARP1-mediated NHEJ repair pathway explains the HTJ format of the initial merges. The findings show that HBV and WHV are immediate inducers of oxidative DNA damage and hijack dsDNA repair to integrate into the hepatocyte genome, and through this mechanism, they may initiate pro-oncogenic processes. Tracking initial integrations may uncover early markers of HCC and help to explain HBV-associated oncogenesis.
Keywords: HBV; dsDNA repair; early hepadnavirus–host DNA integration; hepatocellular carcinoma; oncogenesis; retrotransposons; virus-induced oxidative DNA damage; woodchuck hepatitis virus.
Conflict of interest statement
The author declares no conflict of interest.
Figures


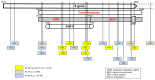

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

