A Novel Role of Connective Tissue Growth Factor in the Regulation of the Epithelial Phenotype
- PMID: 37835529
- PMCID: PMC10571845
- DOI: 10.3390/cancers15194834
A Novel Role of Connective Tissue Growth Factor in the Regulation of the Epithelial Phenotype
Abstract
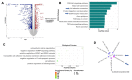
Background: Epithelial-mesenchymal transition (EMT) is a biological process where epithelial cells lose their adhesive properties and gain invasive, metastatic, and mesenchymal properties. Maintaining the balance between the epithelial and mesenchymal stage is essential for tissue homeostasis. Many of the genes promoting mesenchymal transformation have been identified; however, our understanding of the genes responsible for maintaining the epithelial phenotype is limited. Our objective was to identify the genes responsible for maintaining the epithelial phenotype and inhibiting EMT.
Methods: RNA seq was performed using an vitro model of EMT. CTGF expression was determined via qPCR and Western blot analysis. The knockout of CTGF was completed using the CTGF sgRNA CRISPR/CAS9. The tumorigenic potential was determined using NCG mice.
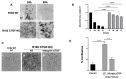
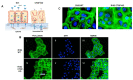
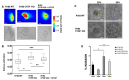
Results: The knockout of CTGF in epithelial ovarian cancer cells leads to the acquisition of functional characteristics associated with the mesenchymal phenotype such as anoikis resistance, cytoskeleton remodeling, increased cell stiffness, and the acquisition of invasion and tumorigenic capacity.
Conclusions: We identified CTGF is an important regulator of the epithelial phenotype, and its loss is associated with the early cellular modifications required for EMT. We describe a novel role for CTGF, regulating cytoskeleton and the extracellular matrix interactions necessary for the conservation of epithelial structure and function. These findings provide a new window into understanding the early stages of mesenchymal transformation.
Keywords: anoikis resistance; cytoskeleton; epithelial–mesenchymal transition; extracellular matrix remodeling; metastasis; ovarian cancer.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



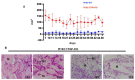
References
-
- American Cancer Society Cancer Facts and Figures 2019. [(accessed on 12 November 2020)]. Available online: https://www.google.com.hk/url?sa=t&rct=j&q=&esrc=s&source=web&cd=&cad=rj....
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

