Genomic monitoring of SARS-CoV-2 variants using sentinel SARI hospital surveillance
- PMID: 37840842
- PMCID: PMC10570899
- DOI: 10.1111/irv.13202
Genomic monitoring of SARS-CoV-2 variants using sentinel SARI hospital surveillance
Abstract
Background: To support the COVID-19 pandemic response, many countries, including Belgium, implemented baseline genomic surveillance (BGS) programs aiming to early detect and characterize new SARS-CoV-2 variants. In parallel, Belgium maintained a sentinel network of six hospitals that samples patients with severe acute respiratory infections (SARI) and integrated SARS-CoV-2 detection within a broader range of respiratory pathogens. We evaluate the ability of the SARI surveillance to monitor general trends and early signals of viral genetic evolution of SARS-CoV-2 and compare it with the BGS as a reference model.
Methods: Nine-hundred twenty-five SARS-CoV-2 positive samples from patients fulfilling the Belgian SARI definition between January 2020 and December 2022 were sequenced using the ARTIC Network amplicon tiling approach on a MinION platform. Weekly variant of concern (VOC) proportions and types were compared to those that were circulating between 2021 and 2022, using 96,251 sequences of the BGS.
Results: SARI surveillance allowed timely detection of the Omicron (BA.1, BA.2, BA.4, and BA.5) and Delta (B.1.617.2) VOCs, with no to 2 weeks delay according to the start of their epidemic growth in the Belgian population. First detection of VOCs B.1.351 and P.1 took longer, but these remained minor in Belgium. Omicron BA.3 was never detected in SARI surveillance. Timeliness could not be evaluated for B.1.1.7, being already major at the start of the study period.
Conclusions: Genomic surveillance of SARS-CoV-2 using SARI sentinel surveillance has proven to accurately reflect VOCs detected in the population and provides a cost-effective solution for long-term genomic monitoring of circulating respiratory viruses.
Keywords: SARI surveillance; SARS‐CoV‐2; genomic surveillance; influenza; pandemic preparedness; respiratory viruses.
© 2023 Sciensano and The Authors. Influenza and Other Respiratory Viruses published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

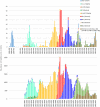
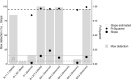
 ) in the national genomic baseline surveillance for each of the different VOCs (Alpha, Beta, Gamma, Delta, Omicron, and its relatives) and the slope (● / ○) and R
2 (◆) values of linear regression curves for each (see the supporting information) were combined to define an emerging variant. A VOC was considered emerging if complying with the following empirical criteria: R
2 > 0.95 (dashed line), slope > 5 (dotted line), and >15% weekly incidence (full line). Based on this definition, Variants B.1.1.7, B.1.617.2, BA.1, BA.2, and BA.5 are considered emerging VOCs, while P.1, BA.3, and BA.4 did not emerge. A slope could not be calculated for VOC B.1.351, as its epidemic growth was not exponential.
) in the national genomic baseline surveillance for each of the different VOCs (Alpha, Beta, Gamma, Delta, Omicron, and its relatives) and the slope (● / ○) and R
2 (◆) values of linear regression curves for each (see the supporting information) were combined to define an emerging variant. A VOC was considered emerging if complying with the following empirical criteria: R
2 > 0.95 (dashed line), slope > 5 (dotted line), and >15% weekly incidence (full line). Based on this definition, Variants B.1.1.7, B.1.617.2, BA.1, BA.2, and BA.5 are considered emerging VOCs, while P.1, BA.3, and BA.4 did not emerge. A slope could not be calculated for VOC B.1.351, as its epidemic growth was not exponential.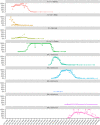
References
-
- World Health Organization . Guidance for surveillance of SARS‐CoV‐2 variants: interim guidance, 9 August 2021. 2021. Accessed October 31, 2022. https://www.who.int/publications/i/item/WHO_2019-nCoV_surveillance_variants
-
- European Centre for Disease Prevention and Control . Guidance for representative and targeted genomic SARS‐CoV‐2 monitoring. 2021. Accessed October 31, 2022. https://www.ecdc.europa.eu/sites/default/files/documents/Guidance-for-re...
MeSH terms
Supplementary concepts
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

