HtrA3: a promising prognostic biomarker and therapeutic target for head and neck squamous cell carcinoma
- PMID: 37842043
- PMCID: PMC10573296
- DOI: 10.7717/peerj.16237
HtrA3: a promising prognostic biomarker and therapeutic target for head and neck squamous cell carcinoma
Abstract
Objective: The dysregulation of the human high-temperature requirement A (HtrA) family of serine proteases is associated with many malignancies. However, there are few reports on HtrAs in head and neck squamous cell carcinoma (HNSCC). The aim of this study was to investigate the expression, prognostic value, and biological functions of HtrAs in HNSCC.
Methods: The RNA-sequencing data and clinical data of HNSCC were downloaded from The Cancer Genome Atlas (TCGA) database. The GSE30784 and GSE31056 datasets from the Gene Expression Omnibus (GEO) database were used for further verification. This study explored the differential expression of HtrAs and assessed their potential impact on the prognosis of HNSCC patients using a survival module. Correlations between clinical characteristics and HtrA expression levels were then explored using a Wilcoxon rank sum test. A Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG), and Gene Set Enrichment Analysis (GSEA) were performed using "clusterProfile" in the R software. A Pearson/Spearman correlation test was applied to analyze the relationship between HtrAs and immune infiltration level/checkpoint genes. Validation of HtrA expression levels were carried out by RT-PCR and western blot in human squamous carcinoma cell lines (Fadu and Cal-27) and human non-tumorigenic bronchial epithelium cells (BEAS-2B). Finally, through cell transfection, CCK-8, Ki-67 immunofluorescence, and flow cytometry assays, the effect of HtrA3 knockdown on the malignant biological behavior of HNSCC cells was explored.
Results: The gene expression levels of HtrAs were significantly upregulated and associated with patient age, TNM stage, clinical stage, and TP53 mutation status in the TCGA-HNSCC cohort. High expressions of HtrA1/3 were associated with shorter overall survival, shorter progress-free interval, and lower disease-specific survival in HNSCC. A nomogram for HtrAs was constructed and validated. HtrA-related genes were significantly enriched in the immune response and cell apoptosis pathway. In addition, the expression of HtrAs showed significant correlations with B cells, M cells, DC cell infiltration, and immune infiltration checkpoint (CD276, TNFRSF14). Validation of HtrA expression was carried out by RT-PCR and western blot. Results of in vitro experiments indicated that HtrA3 gene knockdown inhibits the proliferation of FaDu and Cal-27 cells while concurrently promoting apoptosis.
Conclusions: HtrA3 shows significant potential as both a prognostic marker and a promising therapeutic target for HNSCC, highlighting its relevance and importance in future research and potential clinical applications.
Keywords: Head and neck squamous cell carcinoma; HtrA; Immune infiltration; Immunotherapy; Prognosis.
©2023 Chen et al.
Conflict of interest statement
The authors declare there are no competing interests.
Figures




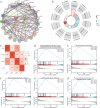



References
-
- Burtness B, Harrington KJ, Greil R, Soulières D, Tahara M, De Castro Jr G, Psyrri A, Basté N, Neupane P, Bratland Å, Fuereder T, Hughes BGM, Mesía R, Ngamphaiboon N, Rordorf T, Wan Ishak WZ, Hong RL, González Mendoza R, Roy A, Zhang Y, Gumuscu B, Cheng JD, Jin F, Rischin D, KEYNOTE-048 Investigators Pembrolizumab alone or with chemotherapy versus cetuximab with chemotherapy for recurrent or metastatic squamous cell carcinoma of the head and neck (KEYNOTE-048): a randomised, open-label, phase 3 study. The Lancet. 2019;394:1915–1928. doi: 10.1016/S0140-6736(19)32591-7. - DOI - PubMed
-
- Chaturvedi AK, Engels EA, Pfeiffer RM, Hernandez BY, Xiao W, Kim E, Jiang B, Goodman MT, Sibug-Saber M, Cozen W, Liu L, Lynch CF, Wentzensen N, Jordan RC, Altekruse S, Anderson WF, Rosenberg PS, Gillison ML. Human papillomavirus and rising oropharyngeal cancer incidence in the United States. Journal of Clinical Oncology. 2011;29:4294–4301. doi: 10.1200/JCO.2011.36.4596. - DOI - PMC - PubMed
-
- Chen YY, Chuang PY, Chen CP, Chiu YH, Lo HF, Cheong ML, Huang JY, Kuo PL, Chen H. Functional antagonism between high temperature requirement protein A (HtrA) family members regulates trophoblast invasion. Journal of Biological Chemistry. 2014;289:22958–22968. doi: 10.1074/jbc.M114.576744. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

