Robust homeostasis of cellular cholesterol is a consequence of endogenous antithetic integral control
- PMID: 37842086
- PMCID: PMC10570530
- DOI: 10.3389/fcell.2023.1244297
Robust homeostasis of cellular cholesterol is a consequence of endogenous antithetic integral control
Abstract
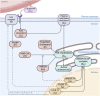
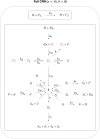
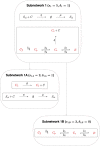
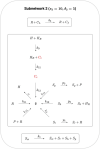
Although cholesterol is essential for cellular viability and proliferation, it is highly toxic in excess. The concentration of cellular cholesterol must therefore be maintained within tight tolerances, and is thought to be subject to a stringent form of homeostasis known as Robust Perfect Adaptation (RPA). While much is known about the cellular signalling interactions involved in cholesterol regulation, the specific chemical reaction network structures that might be responsible for the robust homeostatic regulation of cellular cholesterol have been entirely unclear until now. In particular, the molecular mechanisms responsible for sensing excess whole-cell cholesterol levels have not been identified previously, and no mathematical models to date have been able to capture an integral control implementation that could impose RPA on cellular cholesterol. Here we provide a detailed mathematical description of cholesterol regulation pathways in terms of biochemical reactions, based on an extensive review of experimental and clinical literature. We are able to decompose the associated chemical reaction network structures into several independent subnetworks, one of which is responsible for conferring RPA on several intracellular forms of cholesterol. Remarkably, our analysis reveals that RPA in the cholesterol concentration in the endoplasmic reticulum (ER) is almost certainly due to a well-characterised control strategy known as antithetic integral control which, in this case, involves the high-affinity binding of a multi-molecular transcription factor complex with cholesterol molecules that are excluded from the ER membrane. Our model provides a detailed framework for exploring the necessary biochemical conditions for robust homeostatic control of essential and tightly regulated cellular molecules such as cholesterol.
Keywords: cholesterol homeostasis; integral control; modularity; robust perfect adaptation; signalling network.
Copyright © 2023 Scheepers and Araujo.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
A distributed integral control mechanism for regulation of cholesterol concentration in the human retina.R Soc Open Sci. 2024 Oct 30;11(10):240432. doi: 10.1098/rsos.240432. eCollection 2024 Oct. R Soc Open Sci. 2024. PMID: 39479233 Free PMC article.
-
Antithetic Integral Feedback Ensures Robust Perfect Adaptation in Noisy Biomolecular Networks.Cell Syst. 2016 Jan 27;2(1):15-26. doi: 10.1016/j.cels.2016.01.004. Epub 2016 Jan 27. Cell Syst. 2016. PMID: 27136686
-
Perfect adaptation in biology.Cell Syst. 2021 Jun 16;12(6):509-521. doi: 10.1016/j.cels.2021.05.020. Cell Syst. 2021. PMID: 34139163 Review.
-
Protein-protein complexes can undermine ultrasensitivity-dependent biological adaptation.J R Soc Interface. 2023 Jan;20(198):20220553. doi: 10.1098/rsif.2022.0553. Epub 2023 Jan 4. J R Soc Interface. 2023. PMID: 36596458 Free PMC article.
-
Design Principles Underlying Robust Adaptation of Complex Biochemical Networks.Methods Mol Biol. 2023;2634:3-32. doi: 10.1007/978-1-0716-3008-2_1. Methods Mol Biol. 2023. PMID: 37074572 Free PMC article. Review.
Cited by
-
Identification of tacrolimus-related genes in familial combined hyperlipidemia and development of a diagnostic model using bioinformatics analysis.Heliyon. 2025 Jan 19;11(3):e41705. doi: 10.1016/j.heliyon.2025.e41705. eCollection 2025 Feb 15. Heliyon. 2025. PMID: 39916852 Free PMC article.
-
A distributed integral control mechanism for regulation of cholesterol concentration in the human retina.R Soc Open Sci. 2024 Oct 30;11(10):240432. doi: 10.1098/rsos.240432. eCollection 2024 Oct. R Soc Open Sci. 2024. PMID: 39479233 Free PMC article.
-
Research progress on cholesterol metabolism and tumor therapy.Discov Oncol. 2025 Apr 30;16(1):647. doi: 10.1007/s12672-025-02430-5. Discov Oncol. 2025. PMID: 40307614 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources

