Origins of lineage-specific elements via gene duplication, relocation, and regional rearrangement in Neurospora crassa
- PMID: 37843462
- PMCID: PMC11628664
- DOI: 10.1111/mec.17168
Origins of lineage-specific elements via gene duplication, relocation, and regional rearrangement in Neurospora crassa
Abstract
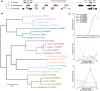
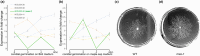
The origin of new genes has long been a central interest of evolutionary biologists. However, their novelty means that they evade reconstruction by the classical tools of evolutionary modelling. This evasion of deep ancestral investigation necessitates intensive study of model species within well-sampled, recently diversified, clades. One such clade is the model genus Neurospora, members of which lack recent gene duplications. Several Neurospora species are comprehensively characterized organisms apt for studying the evolution of lineage-specific genes (LSGs). Using gene synteny, we documented that 78% of Neurospora LSG clusters are located adjacent to the telomeres featuring extensive tracts of non-coding DNA and duplicated genes. Here, we report several instances of LSGs that are likely from regional rearrangements and potentially from gene rebirth. To broadly investigate the functions of LSGs, we assembled transcriptomics data from 68 experimental data points and identified co-regulatory modules using Weighted Gene Correlation Network Analysis, revealing that LSGs are widely but peripherally involved in known regulatory machinery for diverse functions. The ancestral status of the LSG mas-1, a gene with roles in cell-wall integrity and cellular sensitivity to antifungal toxins, was investigated in detail alongside its genomic neighbours, indicating that it arose from an ancient lysophospholipase precursor that is ubiquitous in lineages of the Sordariomycetes. Our discoveries illuminate a "rummage region" in the N. crassa genome that enables the formation of new genes and functions to arise via gene duplication and relocation, followed by fast mutation and recombination facilitated by sequence repeats and unconstrained non-coding sequences.
Keywords: chromosomal rearrangement; de novo origination; fungi; genomics; molecular evolution; orphan gene.
© 2023 The Authors. Molecular Ecology published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Relationship between phylogenetic distribution and genomic features in Neurospora crassa.PLoS One. 2009 Apr 21;4(4):e5286. doi: 10.1371/journal.pone.0005286. PLoS One. 2009. PMID: 19461939 Free PMC article.
-
A natural case of RIP: degeneration of the DNA sequence in an ancestral tandem duplication.Mol Cell Biol. 1989 Oct;9(10):4416-21. doi: 10.1128/mcb.9.10.4416-4421.1989. Mol Cell Biol. 1989. PMID: 2531278 Free PMC article.
-
The genome sequence of the filamentous fungus Neurospora crassa.Nature. 2003 Apr 24;422(6934):859-68. doi: 10.1038/nature01554. Nature. 2003. PMID: 12712197
-
RIP: the evolutionary cost of genome defense.Trends Genet. 2004 Sep;20(9):417-23. doi: 10.1016/j.tig.2004.07.007. Trends Genet. 2004. PMID: 15313550 Review.
-
The Neurospora crassa genome opens up the world of filamentous fungi.Genome Biol. 2003;4(6):217. doi: 10.1186/gb-2003-4-6-217. Epub 2003 May 28. Genome Biol. 2003. PMID: 12801405 Free PMC article. Review.
Cited by
-
The Sordariomycetes: an expanding resource with Big Data for mining in evolutionary genomics and transcriptomics.Front Fungal Biol. 2023 Jun 30;4:1214537. doi: 10.3389/ffunb.2023.1214537. eCollection 2023. Front Fungal Biol. 2023. PMID: 37746130 Free PMC article. Review.
-
Lineage-specific genes are clustered with HET-domain genes and respond to environmental and genetic manipulations regulating reproduction in Neurospora.PLoS Genet. 2023 Nov 7;19(11):e1011019. doi: 10.1371/journal.pgen.1011019. eCollection 2023 Nov. PLoS Genet. 2023. PMID: 37934795 Free PMC article.
References
-
- Basenko, E. Y. , Pulman, J. A. , Shanmugasundram, A. , Harb, O. S. , Crouch, K. , Starns, D. , Warrenfeltz, S. , Aurrecoechea, C. , Stoeckert, C. J., Jr. , Kissinger, J. C. , Roos, D. S. , & Hertz‐Fowler, C. (2018). FungiDB: An integrated bioinformatic resource for fungi and oomycetes. Journal of Fungi (Basel, Switzerland), 4(1), 39. 10.3390/jof4010039 - DOI - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

