From monkeys to humans: observation-basedEMGbrain-computer interface decoders for humans with paralysis
- PMID: 37844567
- PMCID: PMC10618714
- DOI: 10.1088/1741-2552/ad038e
From monkeys to humans: observation-basedEMGbrain-computer interface decoders for humans with paralysis
Abstract
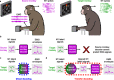
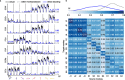
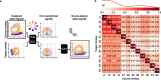
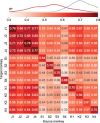
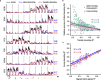
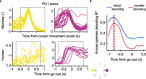
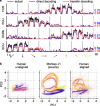
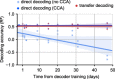
Objective. Intracortical brain-computer interfaces (iBCIs) aim to enable individuals with paralysis to control the movement of virtual limbs and robotic arms. Because patients' paralysis prevents training a direct neural activity to limb movement decoder, most iBCIs rely on 'observation-based' decoding in which the patient watches a moving cursor while mentally envisioning making the movement. However, this reliance on observed target motion for decoder development precludes its application to the prediction of unobservable motor output like muscle activity. Here, we ask whether recordings of muscle activity from a surrogate individual performing the same movement as the iBCI patient can be used as target for an iBCI decoder.Approach. We test two possible approaches, each using data from a human iBCI user and a monkey, both performing similar motor actions. In one approach, we trained a decoder to predict the electromyographic (EMG) activity of a monkey from neural signals recorded from a human. We then contrast this to a second approach, based on the hypothesis that the low-dimensional 'latent' neural representations of motor behavior, known to be preserved across time for a given behavior, might also be preserved across individuals. We 'transferred' an EMG decoder trained solely on monkey data to the human iBCI user after using Canonical Correlation Analysis to align the human latent signals to those of the monkey.Main results. We found that both direct and transfer decoding approaches allowed accurate EMG predictions between two monkeys and from a monkey to a human.Significance. Our findings suggest that these latent representations of behavior are consistent across animals and even primate species. These methods are an important initial step in the development of iBCI decoders that generate EMG predictions that could serve as signals for a biomimetic decoder controlling motion and impedance of a prosthetic arm, or even muscle force directly through functional electrical stimulation.
Keywords: EMG decoding; brain computer interface; monkey; paralyzed human; transfer learning.
Creative Commons Attribution license.
Conflict of interest statement
The authors declare no competing interest.
Figures

References
Publication types
MeSH terms
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
