Brucella MucR acts as an H-NS-like protein to silence virulence genes and structure the nucleoid
- PMID: 37847580
- PMCID: PMC10746212
- DOI: 10.1128/mbio.02201-23
Brucella MucR acts as an H-NS-like protein to silence virulence genes and structure the nucleoid
Abstract
Histone-like nucleoid structuring (H-NS) and H-NS-like proteins coordinate host-associated behaviors in many pathogenic bacteria, often through forming silencer/counter-silencer pairs with signal-responsive transcriptional activators to tightly control gene expression. Brucella and related bacteria do not encode H-NS or homologs of known H-NS-like proteins, and it is unclear if they have other proteins that perform analogous functions during pathogenesis. In this work, we provide compelling evidence for the role of MucR as a novel H-NS-like protein in Brucella. We show that MucR possesses many of the known functions attributed to H-NS and H-NS-like proteins, including the formation of silencer/counter-silencer pairs to control virulence gene expression and global structuring of the nucleoid. These results uncover a new role for MucR as a nucleoid structuring protein and support the importance of temporal control of gene expression in Brucella and related bacteria.
Keywords: Brucella; ChIP-seq; H-NS; H-NS-like; Hi-C; MucR; counter-silencer; nucleoid structuring; virulence.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


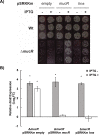
References
-
- Altamirano-Silva P, Meza-Torres J, Castillo-Zeledón A, Ruiz-Villalobos N, Zuñiga-Pereira AM, Chacón-Díaz C, Moreno E, Guzmán-Verri C, Chaves-Olarte E. 2018. Brucella abortus senses the intracellular environment through the BvrR/BvrS two-component system, which allows B. abortus to adapt to its replicative niche. Infect Immun 86:e00713-17. doi:10.1128/IAI.00713-17 - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
- R01 GM141242/GM/NIGMS NIH HHS/United States
- R01 GM143182/GM/NIGMS NIH HHS/United States
- R21 AI141438/AI/NIAID NIH HHS/United States
- AI141438/HHS | NIH | National Institute of Allergy and Infectious Diseases (NIAID)
- GM141242, GM143182/HHS | NIH | National Institute of General Medical Sciences (NIGMS)
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
