Endotype Characterization Reveals Mechanistic Differences Across Brain Regions in Sporadic Alzheimer's Disease
- PMID: 37849634
- PMCID: PMC10578327
- DOI: 10.3233/ADR-220098
Endotype Characterization Reveals Mechanistic Differences Across Brain Regions in Sporadic Alzheimer's Disease
Abstract
Background: While Alzheimer's disease (AD) pathology is associated with altered brain structure, it is not clear whether gene expression changes mirror the onset and evolution of pathology in distinct brain regions. Deciphering the mechanisms which cause the differential manifestation of the disease across different regions has the potential to help early diagnosis.
Objective: We aimed to identify common and unique endotypes and their regulation in tangle-free neurons in sporadic AD (SAD) across six brain regions: entorhinal cortex (EC), hippocampus (HC), medial temporal gyrus (MTG), posterior cingulate (PC), superior frontal gyrus (SFG), and visual cortex (VCX).
Methods: To decipher the states of tangle-free neurons across different brain regions in human subjects afflicted with AD, we performed analysis of the neural transcriptome. We explored changes in differential gene expression, functional and transcription factor target enrichment, and co-expression gene module detection analysis to discern disease-state transcriptomic variances and characterize endotypes. Additionally, we compared our results to tangled AD neuron microarray-based study and the Allen Brain Atlas.
Results: We identified impaired neuron function in EC, MTG, PC, and VCX resulting from REST activation and reversal of mature neurons to a precursor-like state in EC, MTG, and SFG linked to SOX2 activation. Additionally, decreased neuron function and increased dedifferentiation were linked to the activation of SUZ12. Energetic deficit connected to NRF1 inactivation was found in HC, PC, and VCX.
Conclusions: Our findings suggest that SAD manifestation varies in scale and severity in different brain regions. We identify endotypes, such as energetic shortfalls, impaired neuronal function, and dedifferentiation.
Keywords: Alzheimer’s disease; NRF1; REST; SOX2; SUZ12; dedifferentiation; endotype; energetics; sporadic Alzheimer’s disease; transcriptome.
© 2023 – The authors. Published by IOS Press.
Conflict of interest statement
The authors have no conflict of interest to report.
Figures
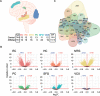




References
-
- Hendrie HC (1998) Epidemiology of dementia and Alzheimer’s disease. Am J Geriatr Psychiatry 6, S3–18. - PubMed
-
- Tang M, Ryman DC, McDade E, Jasielec MS, Buckles VD, Cairns NJ, Fagan AM, Goate A, Marcus DS, Xiong C, Allegri RF, Chhatwal JP, Danek A, Farlow MR, Fox NC, Ghetti B, Graff-Radford NR, Laske C, Martins RN, Masters CL, Mayeux RP, Ringman JM, Rossor MN, Salloway SP, Schofield PR, Morris JC, Bateman RJ; Dominantly Inherited Alzheimer Network (DIAN) (2016) Neurological manifestations of autosomal dominant familial Alzheimer’s disease: A comparison of the published literature with the Dominantly Inherited Alzheimer Network observational study (DIAN-OBS). Lancet Neurol 15, 1317–1325. Erratum in: Lancet Neurol. 2017 Jan;16(1):24. - PMC - PubMed
-
- Shepherd C, McCann H, Halliday GM (2009) Variations in the neuropathology of familial Alzheimer’s disease. Acta Neuropathol (Berl) 118, 37–52. - PubMed
-
- Roher AE, Maarouf CL, Kokjohn TA (2016) Familial presenilin mutations and sporadic Alzheimer’s disease pathology: Is the assumption of biochemical equivalence justified? J Alzheimers Dis 50, 645–658. - PubMed

