Revolutionizing antiretroviral therapy for human immunodeficiency virus/AIDS: A computational approach using molecular docking, virtual screening, and 3D pharmacophore building to address therapeutic failure and propose highly effective candidates
- PMID: 37850462
- PMCID: PMC10585989
- DOI: 10.1177/03946320231207514
Revolutionizing antiretroviral therapy for human immunodeficiency virus/AIDS: A computational approach using molecular docking, virtual screening, and 3D pharmacophore building to address therapeutic failure and propose highly effective candidates
Abstract
Objectives: In the context of human immunodeficiency virus (HIV) treatment, the emergence of therapeutic failures with existing antiretroviral drugs presents a significant challenge. This study aims to employ advanced molecular modeling techniques to identify potential alternatives to current antiretroviral agents.
Methods: The study focuses on three essential classes of antiretroviral drugs: nucleoside reverse transcriptase inhibitors (NRTIs), non-nucleoside reverse transcriptase inhibitors (NNRTIs), and protease inhibitors (PIs). Computational analyses were performed on a database of 3,343,652 chemical molecules to evaluate their binding affinities, pharmacokinetic properties, and interactions with viral reverse transcriptase and protease enzymes. Molecular docking, virtual screening, and 3D pharmacophore modeling were utilized to identify promising candidates.
Results: Molecular docking revealed compounds with high binding energies and strong interactions at the active sites of target enzymes. Virtual screening narrowed down potential candidates with favorable pharmacological profiles. 3D pharmacophore modeling identified crucial structural features for effective binding. Overall, two molecules for class 1, 7 molecules for class 2, and 2 molecules for class 3 were selected. These compounds exhibited robust binding affinities, interactions with target enzymes, and improved pharmacokinetic properties, showing promise for more effective HIV treatments in cases of therapeutic failures.
Conclusion: The combination of molecular docking, virtual screening, and 3D pharmacophore modeling yielded lead compounds that hold potential for addressing HIV therapeutic failures. Further experimental investigations are essential to validate the efficacy and safety of these compounds, with the ultimate goal of advancing toward clinical applications in HIV management.
Keywords: antiretroviral; computational screening; human immunodeficiency virus-failure of treatment; pharmacophore model.
Conflict of interest statement
Declaration of conflicting interestsThe author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Figures

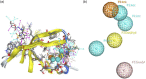

Similar articles
-
Proposal of pharmacophore model for HIV reverse transcriptase inhibitors: Combined mutational effect analysis, molecular dynamics, molecular docking and pharmacophore modeling study.Int J Immunopathol Pharmacol. 2024 Jan-Dec;38:3946320241231465. doi: 10.1177/03946320241231465. Int J Immunopathol Pharmacol. 2024. PMID: 38296818 Free PMC article.
-
A Combined Approach of Pharmacophore Modeling, QSAR Study, Molecular Docking and In silico ADME/Tox Prediction of 4-Arylthio & 4-Aryloxy-3- Iodopyridine-2(1H)-one Analogs to Identify Potential Reverse Transcriptase Inhibitor: Anti-HIV Agents.Med Chem. 2022;18(1):51-87. doi: 10.2174/1573406417666201214100822. Med Chem. 2022. PMID: 33319692
-
Current insights and molecular docking studies of HIV-1 reverse transcriptase inhibitors.Chem Biol Drug Des. 2024 Jan;103(1):e14372. doi: 10.1111/cbdd.14372. Epub 2023 Oct 10. Chem Biol Drug Des. 2024. PMID: 37817296 Review.
-
A Step Towards Optimization of Amide-Linked Coumarin Pharmacophore: As an Anti-HIV Agent.Curr HIV Res. 2024;22(5):279-289. doi: 10.2174/011570162X308550240821074309. Curr HIV Res. 2024. PMID: 39317999
-
The development of anti-HIV-1 drugs.Yao Xue Xue Bao. 2010 Feb;45(2):165-76. Yao Xue Xue Bao. 2010. PMID: 21348415 Review.
Cited by
-
Syk protein inhibitors treatment for the allergic symptoms associated with hyper immunoglobulin E syndromes: A focused on a computational approach.Int J Immunopathol Pharmacol. 2024 Jan-Dec;38:3946320241282030. doi: 10.1177/03946320241282030. Int J Immunopathol Pharmacol. 2024. PMID: 39241232 Free PMC article.
References
-
- Sluis-Cremer N. (2021) Retroviral reverse transcriptase: structure, function and inhibition. The Enzymes 50: 179–194. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

