Long 3'UTRs predispose neurons to inflammation by promoting immunostimulatory double-stranded RNA formation
- PMID: 37862432
- PMCID: PMC11056275
- DOI: 10.1126/sciimmunol.adg2979
Long 3'UTRs predispose neurons to inflammation by promoting immunostimulatory double-stranded RNA formation
Abstract
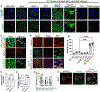
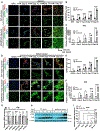
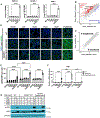
Loss of RNA homeostasis underlies numerous neurodegenerative and neuroinflammatory diseases. However, the molecular mechanisms that trigger neuroinflammation are poorly understood. Viral double-stranded RNA (dsRNA) triggers innate immune responses when sensed by host pattern recognition receptors (PRRs) present in all cell types. Here, we report that human neurons intrinsically carry exceptionally high levels of immunostimulatory dsRNAs and identify long 3'UTRs as giving rise to neuronal dsRNA structures. We found that the neuron-enriched ELAVL family of genes (ELAVL2, ELAVL3, and ELAVL4) can increase (i) 3'UTR length, (ii) dsRNA load, and (iii) activation of dsRNA-sensing PRRs such as MDA5, PKR, and TLR3. In wild-type neurons, neuronal dsRNAs signaled through PRRs to induce tonic production of the antiviral type I interferon. Depleting ELAVL2 in WT neurons led to global shortening of 3'UTR length, reduced immunostimulatory dsRNA levels, and rendered WT neurons susceptible to herpes simplex virus and Zika virus infection. Neurons deficient in ADAR1, a dsRNA-editing enzyme mutated in the neuroinflammatory disorder Aicardi-Goutières syndrome, exhibited intolerably high levels of dsRNA that triggered PRR-mediated toxic inflammation and neuronal death. Depleting ELAVL2 in ADAR1 knockout neurons led to prolonged neuron survival by reducing immunostimulatory dsRNA levels. In summary, neurons are specialized cells where PRRs constantly sense "self" dsRNAs to preemptively induce protective antiviral immunity, but maintaining RNA homeostasis is paramount to prevent pathological neuroinflammation.
Figures

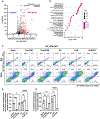


References
-
- Zu T, Guo S, Bardhi O, Ryskamp DA, Li J, Tusi SK, Engelbrecht A, Kiippel K, Chakrabarty P, Nguyen L, Goide TE, Sonenberg N, Ranum LPW, Metformin inhibits RAN translation through PKR pathway and mitigates disease in C9orf72 ALS/FTD mice. Proc. Natl. Acad. Sci. U.S.A 117, 18591–18599 (2020). - PMC - PubMed
-
- Rodriguez S,Sahin A,Schrank BR,Ai-Lawati H,Costantino i.,Benz E,Fard D,Albers AD,Cao L,Gomez AC,Evans K,Ratti E,Cudkowicz M,Frosch MP,Talkowski M,Sorger PK,Hyman BT,Albers MW,Genome-encoded cytoplasmic double-stranded RNAs, found in C9ORF72 ALS-FTD brain, propagate neuronal loss. Sci. Transl. Med 13, eaaz4699 (2021). - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases