Fungal microbiota sustains lasting immune activation of neutrophils and their progenitors in severe COVID-19
- PMID: 37872315
- PMCID: PMC10805066
- DOI: 10.1038/s41590-023-01637-4
Fungal microbiota sustains lasting immune activation of neutrophils and their progenitors in severe COVID-19
Abstract
Gastrointestinal fungal dysbiosis is a hallmark of several diseases marked by systemic immune activation. Whether persistent pathobiont colonization during immune alterations and impaired gut barrier function has a durable impact on host immunity is unknown. We found that elevated levels of Candida albicans immunoglobulin G (IgG) antibodies marked patients with severe COVID-19 (sCOVID-19) who had intestinal Candida overgrowth, mycobiota dysbiosis and systemic neutrophilia. Analysis of hematopoietic stem cell progenitors in sCOVID-19 revealed transcriptional changes in antifungal immunity pathways and reprogramming of granulocyte myeloid progenitors (GMPs) for up to a year. Mice colonized with C. albicans patient isolates experienced increased lung neutrophilia and pulmonary NETosis during severe acute respiratory syndrome coronavirus-2 infection, which were partially resolved with antifungal treatment or by interleukin-6 receptor blockade. sCOVID-19 patients treated with tocilizumab experienced sustained reductions in C. albicans IgG antibodies titers and GMP transcriptional changes. These findings suggest that gut fungal pathobionts may contribute to immune activation during inflammatory diseases, offering potential mycobiota-immune therapeutic strategies for sCOVID-19 with prolonged symptoms.
© 2023. The Author(s), under exclusive licence to Springer Nature America, Inc.
Conflict of interest statement
Declaration of Interests
The AGS laboratory has received research support from GSK, Pfizer, Senhwa Biosciences, Kenall Manufacturing, Blade Therapeutics, Avimex, Johnson & Johnson, Dynavax, 7Hills Pharma, Pharmamar, ImmunityBio, Accurius, Nanocomposix, Hexamer, N-fold LLC, Model Medicines, Atea Pharma, Applied Biological Laboratories and Merck, outside of the reported work. A.G.-S. has consulting agreements for the following companies involving cash and/or stock: Castlevax, Amovir, Vivaldi Biosciences, Contrafect, 7Hills Pharma, Avimex, Pagoda, Accurius, Esperovax, Farmak, Applied Biological Laboratories, Pharmamar, CureLab Oncology, CureLab Veterinary, Synairgen, Paratus, Pfizer and Prosetta, outside of the reported work. A.G.-S. has been an invited speaker in meeting events organized by Seqirus, Janssen, Abbott and Astrazeneca. A.G.-S. is inventor on patents and patent applications on the use of antivirals and vaccines for the treatment and prevention of virus infections and cancer, owned by the Icahn School of Medicine at Mount Sinai, New York, outside of the reported work. M Schotsaert has received unrelated research funding in sponsored research agreements from ArgenX BV, Moderna, 7Hills Pharma and Phio Pharmaceuticals which has no competing interest with this work. The other authors declare no competing interests related to this study.
Figures







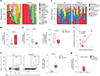



References
-
- COVID-19 Dashboard. the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University (JHU).
Methods REFERENCES
MeSH terms
Substances
Grants and funding
- K08 MH130773/MH/NIMH NIH HHS/United States
- R01 AI143861/AI/NIAID NIH HHS/United States
- R01 DK121977/DK/NIDDK NIH HHS/United States
- R01 DK130425/DK/NIDDK NIH HHS/United States
- R56 AI137157/AI/NIAID NIH HHS/United States
- R01 AI160706/AI/NIAID NIH HHS/United States
- R01 DK113136/DK/NIDDK NIH HHS/United States
- U19 AI168631/AI/NIAID NIH HHS/United States
- P30 CA016087/CA/NCI NIH HHS/United States
- U19 AI142733/AI/NIAID NIH HHS/United States
- R01 AI163007/AI/NIAID NIH HHS/United States
- U19 AI135972/AI/NIAID NIH HHS/United States
- 75N93021C00014/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Research Materials

