Temporal and structural patterns of hepatitis B virus integrations in hepatocellular carcinoma
- PMID: 37877809
- PMCID: PMC11131385
- DOI: 10.1002/jmv.29187
Temporal and structural patterns of hepatitis B virus integrations in hepatocellular carcinoma
Abstract
Chronic infection of hepatitis B virus (HBV) is the major cause of hepatocellular carcinoma (HCC). Notably, 90% of HBV-positive HCC cases exhibit detectable HBV integrations, hinting at the potential early entanglement of these viral integrations in tumorigenesis and their subsequent oncogenic implications. Nevertheless, the precise chronology of integration events during HCC tumorigenesis, alongside their sequential structural patterns, has remained elusive thus far. In this study, we applied whole-genome sequencing to multiple biopsies extracted from six HBV-positive HCC cases. Through this approach, we identified point mutations and viral integrations, offering a blueprint for the intricate tumor phylogeny of these samples. The emergent narrative paints a rich tapestry of diverse evolutionary trajectories characterizing the analyzed tumors. We uncovered oncogenic integration events in some samples that appear to happen before and during the initiation stage of tumor development based on their locations in reconstituted trajectories. Furthermore, we conducted additional long-read sequencing of selected samples and unveiled integration-bridged chromosome rearrangements and tandem repeats of the HBV sequence within integrations. In summary, this study revealed premalignant oncogenic and sequential complex integrations and highlighted the contributions of HBV integrations to HCC development and genome instability.
Keywords: chromosomal translocation; clonal evolution; hepatocellular carcinoma; intratumor heterogeneity; multiregion sequencing; viral integration.
© 2023 The Authors. Journal of Medical Virology Published by Wiley Periodicals LLC.
Conflict of interest statement
Conflict of Interest statement
The authors declare that there is no conflict of interest.
Figures


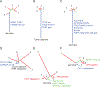

References
-
- Levrero M and Zucman-Rossi J, Mechanisms of HBV-induced hepatocellular carcinoma. J Hepatol, 2016. 64(1 Suppl): p. S84–s101. - PubMed

