Recessive COL17A1 Mutations and a Dominant LAMB3 Mutation Cause Hypoplastic Amelogenesis Imperfecta
- PMID: 37888105
- PMCID: PMC10608278
- DOI: 10.3390/jpm13101494
Recessive COL17A1 Mutations and a Dominant LAMB3 Mutation Cause Hypoplastic Amelogenesis Imperfecta
Abstract
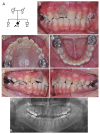
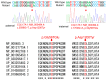
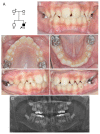
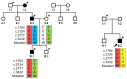
Hereditary conditions that affect tooth enamel in quantity and/or quality are called amelogenesis imperfecta (AI). AI can occur as an isolated condition or as a symptom of a syndrome. An OMIM search with the term "AI" yielded 79 result entries. Mutations in the same gene cause syndromic or non-syndromic AI, depending on the nature of the mutations. In this study, we recruited two AI families and performed mutational analysis using whole-exome sequencing. The proband of family 1, with hypoplastic pitted AI and mild localized atopic dermatitis, had compound heterozygous COL17A1 mutations (paternal NM_000494.4: c.3598G>T, p.Asp1200Tyr and maternal c.1700G>A, p.Gly567Glu). The proband of family 2, with hypoplastic pitted AI and Jervell and Lange-Nielsen syndrome, had a recurrent LAMB3 mutation (NM_000228.3: c.3463_3475del, p.(Glu1155Thrfs*51)) in addition to compound heterozygous mutations in the KCNQ1 gene.
Keywords: COL17A1; LAMB3; enamel defects; hereditary; mutation.
Conflict of interest statement
The authors declared no potential conflict of interest with respect to the research, authorship, and/or publication of this article.
Figures

Similar articles
-
Novel ENAM and LAMB3 mutations in Chinese families with hypoplastic amelogenesis imperfecta.PLoS One. 2015 Mar 13;10(3):e0116514. doi: 10.1371/journal.pone.0116514. eCollection 2015. PLoS One. 2015. PMID: 25769099 Free PMC article.
-
LAMB3 mutations causing autosomal-dominant amelogenesis imperfecta.J Dent Res. 2013 Oct;92(10):899-904. doi: 10.1177/0022034513502054. Epub 2013 Aug 19. J Dent Res. 2013. PMID: 23958762 Free PMC article.
-
Phenotype and Variant Spectrum in the LAMB3 Form of Amelogenesis Imperfecta.J Dent Res. 2019 Jun;98(6):698-704. doi: 10.1177/0022034519835205. Epub 2019 Mar 24. J Dent Res. 2019. PMID: 30905256 Free PMC article.
-
Molecular-based phenotype variations in amelogenesis imperfecta.Oral Dis. 2023 Sep;29(6):2334-2365. doi: 10.1111/odi.14599. Epub 2023 May 8. Oral Dis. 2023. PMID: 37154292 Review.
-
Amelogenesis Imperfecta; Genes, Proteins, and Pathways.Front Physiol. 2017 Jun 26;8:435. doi: 10.3389/fphys.2017.00435. eCollection 2017. Front Physiol. 2017. PMID: 28694781 Free PMC article. Review.
Cited by
-
Unveiling Collagen's Role in Breast Cancer: Insights into Expression Patterns, Functions and Clinical Implications.Int J Gen Med. 2024 May 2;17:1773-1787. doi: 10.2147/IJGM.S463649. eCollection 2024. Int J Gen Med. 2024. PMID: 38711825 Free PMC article. Review.
References
Grants and funding
LinkOut - more resources
Full Text Sources

