An evolution strategy approach for the balanced minimum evolution problem
- PMID: 37889263
- PMCID: PMC10641104
- DOI: 10.1093/bioinformatics/btad660
An evolution strategy approach for the balanced minimum evolution problem
Abstract
Motivation: The Balanced Minimum Evolution (BME) is a powerful distance based phylogenetic estimation model introduced by Desper and Gascuel and nowadays implemented in popular tools for phylogenetic analyses. It was proven to be computationally less demanding than more sophisticated estimation methods, e.g. maximum likelihood or Bayesian inference while preserving the statistical consistency and the ability to run with almost any kind of data for which a dissimilarity measure is available. BME can be stated in terms of a nonlinear non-convex combinatorial optimization problem, usually referred to as the Balanced Minimum Evolution Problem (BMEP). Currently, the state-of-the-art among approximate methods for the BMEP is represented by FastME (version 2.0), a software which implements several deterministic phylogenetic construction heuristics combined with a local search on specific neighbourhoods derived by classical topological tree rearrangements. These combinations, however, may not guarantee convergence to close-to-optimal solutions to the problem due to the lack of solution space exploration, a phenomenon which is exacerbated when tackling molecular datasets characterized by a large number of taxa.
Results: To overcome such convergence issues, in this article, we propose a novel metaheuristic, named PhyloES, which exploits the combination of an exploration phase based on Evolution Strategies, a special type of evolutionary algorithm, with a refinement phase based on two local search algorithms. Extensive computational experiments show that PhyloES consistently outperforms FastME, especially when tackling larger datasets, providing solutions characterized by a shorter tree length but also significantly different from the topological perspective.
Availability and implementation: The software and the data are available at https://github.com/andygaspar/PHYLOES.
© The Author(s) 2023. Published by Oxford University Press.
Conflict of interest statement
None declared.
Figures
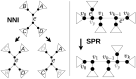


References
-
- Bäck T. Evolutionary Algorithms in Theory and Practice: Evolution Strategies, Evolutionary Programming, Genetic Algorithms. Oxford, UK: Oxford University Press, 1996.
-
- Bartoli A, De Lorenzo A, Medvet E. et al. Multi-level diversity promotion strategies for grammar-guided genetic programming. Applied Soft Computing 2019;83:105599.
-
- Bordewich M, Gascuel O, Huber KT. et al. Consistency of topological moves based on the balanced minimum evolution principle of phylogenetic inference. IEEE/ACM Trans Comput Biol Bioinform 2009;6:110–7. - PubMed
-
- Brauer MJ, Holder MT, Dries LA. et al. Genetic algorithms and parallel processing in maximum-likelihood phylogeny inference. Mol Biol Evol 2002;19:1717–26. - PubMed

