Chemical Insights into Oxidative and Nitrative Modifications of DNA
- PMID: 37894920
- PMCID: PMC10607741
- DOI: 10.3390/ijms242015240
Chemical Insights into Oxidative and Nitrative Modifications of DNA
Abstract
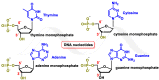
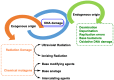
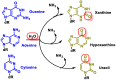
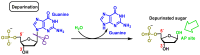
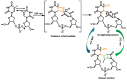
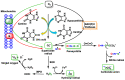
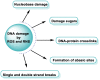
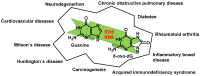
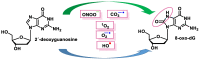
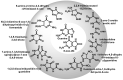
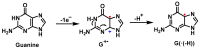
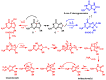
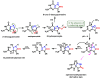
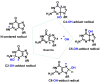
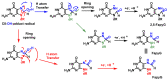
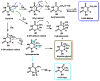
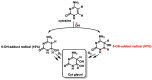
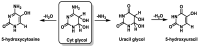
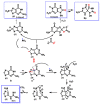
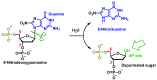
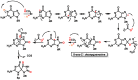
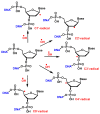
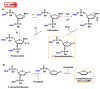
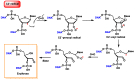
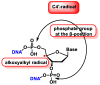
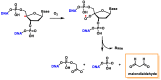
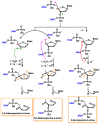
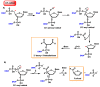
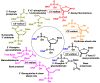
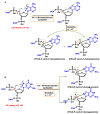
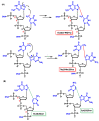
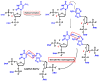
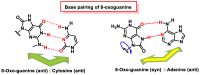
This review focuses on DNA damage caused by a variety of oxidizing, alkylating, and nitrating species, and it may play an important role in the pathophysiology of inflammation, cancer, and degenerative diseases. Infection and chronic inflammation have been recognized as important factors in carcinogenesis. Under inflammatory conditions, reactive oxygen species (ROS) and reactive nitrogen species (RNS) are generated from inflammatory and epithelial cells, and result in the formation of oxidative and nitrative DNA lesions, such as 8-oxo-7,8-dihydro-2'-deoxyguanosine (8-oxodG) and 8-nitroguanine. Cellular DNA is continuously exposed to a very high level of genotoxic stress caused by physical, chemical, and biological agents, with an estimated 10,000 modifications occurring every hour in the genetic material of each of our cells. This review highlights recent developments in the chemical biology and toxicology of 2'-deoxyribose oxidation products in DNA.
Keywords: 2′-deoxyribose oxidation products; DNA; inflammation; oxidative and nitrative DNA damage.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures
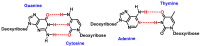


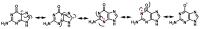
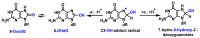

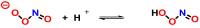
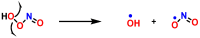
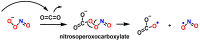


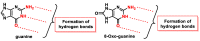

Similar articles
-
Nitrative and oxidative DNA damage in infection-related carcinogenesis in relation to cancer stem cells.Genes Environ. 2017 Jan 1;38:26. doi: 10.1186/s41021-016-0055-7. eCollection 2016. Genes Environ. 2017. PMID: 28050219 Free PMC article. Review.
-
DNA damage in inflammation-related carcinogenesis and cancer stem cells.Oxid Med Cell Longev. 2013;2013:387014. doi: 10.1155/2013/387014. Epub 2013 Dec 5. Oxid Med Cell Longev. 2013. PMID: 24382987 Free PMC article. Review.
-
Crosstalk between DNA Damage and Inflammation in the Multiple Steps of Carcinogenesis.Int J Mol Sci. 2017 Aug 19;18(8):1808. doi: 10.3390/ijms18081808. Int J Mol Sci. 2017. PMID: 28825631 Free PMC article. Review.
-
Cancer risk and oxidative DNA damage in man.J Mol Med (Berl). 1996 Jun;74(6):297-312. doi: 10.1007/BF00207507. J Mol Med (Berl). 1996. PMID: 8862511 Review.
-
Oxidative and nitrative DNA damage in animals and patients with inflammatory diseases in relation to inflammation-related carcinogenesis.Biol Chem. 2006 Apr;387(4):365-72. doi: 10.1515/BC.2006.049. Biol Chem. 2006. PMID: 16606333 Review.
Cited by
-
Generation of cellular reactive oxygen and nitrogen species by exposure to ultraviolet radiation.Biophys Rev. 2025 Mar 14;17(2):547-560. doi: 10.1007/s12551-025-01298-7. eCollection 2025 Apr. Biophys Rev. 2025. PMID: 40376412 Review.
-
Deprotonation of 8-Oxo-7,8-dihydroadenine Radical Cation in Free and Encumbered Context: A Theoretical Study.ACS Omega. 2024 Dec 13;9(51):50730-50741. doi: 10.1021/acsomega.4c08956. eCollection 2024 Dec 24. ACS Omega. 2024. PMID: 39741838 Free PMC article.
-
Methods and applications of genome-wide profiling of DNA damage and rare mutations.Nat Rev Genet. 2024 Dec;25(12):846-863. doi: 10.1038/s41576-024-00748-4. Epub 2024 Jun 25. Nat Rev Genet. 2024. PMID: 38918545 Free PMC article. Review.
-
o8G-modified circKIAA1797 promotes lung cancer development by inhibiting cuproptosis.J Exp Clin Cancer Res. 2025 Apr 2;44(1):110. doi: 10.1186/s13046-025-03365-z. J Exp Clin Cancer Res. 2025. PMID: 40176113 Free PMC article.
-
Reactive Nitrogen Species and Fibrinogen: Exploring the Effects of Nitration on Blood Clots.Antioxidants (Basel). 2025 Jul 4;14(7):825. doi: 10.3390/antiox14070825. Antioxidants (Basel). 2025. PMID: 40722929 Free PMC article. Review.
References
-
- Alberts B. Molecular Biology of the Cell. WW Norton & Company; New York, NY, USA: 2017.
Publication types
MeSH terms
Substances
Grants and funding
- ProID2020010134/Agencia Canaria de Investigación, Innovación y Sociedad de la Información
- Project 2019SP43/Fundación CajaCanarias
- PLEC2022-009507/State Plan for Scientific, Technical Research and Innovation 2021-2023 from the Spanish Ministry of Science and Innovation
- PID2019-105838RB-C31/Spanish Ministry of Economy and Competitiveness
LinkOut - more resources
Full Text Sources
Medical

