Transcriptional Analysis of lncRNA and Target Genes Induced by Influenza A Virus Infection in MDCK Cells
- PMID: 37896995
- PMCID: PMC10610897
- DOI: 10.3390/vaccines11101593
Transcriptional Analysis of lncRNA and Target Genes Induced by Influenza A Virus Infection in MDCK Cells
Abstract
Background: The MDCK cell line is the primary cell line used for influenza vaccine production. Using genetic engineering technology to change the expression and activity of genes that regulate virus proliferation to obtain high-yield vaccine cell lines has attracted increasing attention. A comprehensive understanding of the key genes, targets, and molecular mechanisms of viral regulation in cells is critical to achieving this goal, yet the post-transcriptional regulation mechanism involved in virus proliferation-particularly the effect of lncRNA on influenza virus proliferation-is still poorly understood. Therefore, this study used high-throughput RNA-seq technology to identify H1N1 infection-induced lncRNA and mRNA expression changes in MDCK cells and explore the regulatory relationship between these crucial lncRNAs and their target genes.
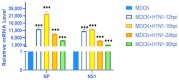
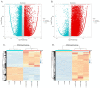
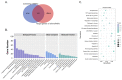
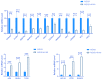
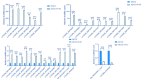
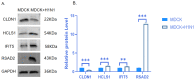
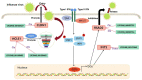
Results: In response to H1N1 infection in MDCK cells 16 h post-infection (hpi) relative to uninfected controls, we used multiple gene function annotation databases and initially identified 31,501 significantly differentially expressed (DE) genes and 39,920 DE lncRNAs (|log2FC| > 1, p < 0.05). Among these, 102 lncRNAs and 577 mRNAs exhibited predicted correlations with viral response mechanisms. Based on the magnitude of significant expression differences, related research, and RT-qPCR expression validation at the transcriptional level, we further focused on 18 DE mRNAs and 32 DE lncRNAs. Among these, the differential expression of the genes RSAD2, CLDN1, HCLS1, and IFIT5 in response to influenza virus infection was further verified at the protein level using Western blot technology, which showed results consistent with the RNA-seq and RT-qPCR findings. We then developed a potential molecular regulatory network between these four genes and their six predicted lncRNAs.
Conclusions: The results of this study will contribute to a more comprehensive understanding of the molecular mechanism of host cell non-coding RNA-mediated regulation of influenza virus replication. These results may also identify methods for screening target genes in the development of genetically engineered cell lines capable of high-yield artificial vaccine production.
Keywords: IAV; MDCK; influenza vaccine; lncRNAs; transcriptomics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
RSAD2 Is an Effective Target for High-Yield Vaccine Production in MDCK Cells.Viruses. 2022 Nov 21;14(11):2587. doi: 10.3390/v14112587. Viruses. 2022. PMID: 36423196 Free PMC article.
-
Absolute quantification of viral proteins during single-round replication of MDCK suspension cells.J Proteomics. 2022 May 15;259:104544. doi: 10.1016/j.jprot.2022.104544. Epub 2022 Mar 1. J Proteomics. 2022. PMID: 35240312
-
Transcriptome-wide 5-methylcytosine modification profiling of long non-coding RNAs in A549 cells infected with H1N1 influenza A virus.BMC Genomics. 2023 Jun 12;24(1):316. doi: 10.1186/s12864-023-09432-z. BMC Genomics. 2023. PMID: 37308824 Free PMC article.
-
Roles of lncRNAs in influenza virus infection.Emerg Microbes Infect. 2020 Dec;9(1):1407-1414. doi: 10.1080/22221751.2020.1778429. Emerg Microbes Infect. 2020. PMID: 32543285 Free PMC article. Review.
-
Host Non-Coding RNA Regulates Influenza A Virus Replication.Viruses. 2021 Dec 29;14(1):51. doi: 10.3390/v14010051. Viruses. 2021. PMID: 35062254 Free PMC article. Review.
Cited by
-
Production, Passaging Stability, and Histological Analysis of Madin-Darby Canine Kidney Cells Cultured in a Low-Serum Medium.Vaccines (Basel). 2024 Aug 30;12(9):991. doi: 10.3390/vaccines12090991. Vaccines (Basel). 2024. PMID: 39340023 Free PMC article.
References
-
- Iuliano A.D., Roguski K.M., Chang H.H., Muscatello D.J., Palekar R., Tempia S., Cohen C., Gran J.M., Schanzer D., Cowling B.J., et al. Estimates of global seasonal influenza-associated respiratory mortality: A modelling study. Lancet. 2018;391:1285–1300. doi: 10.1016/S0140-6736(17)33293-2. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

