Metagenomics analysis reveals presence of the Merida-like virus in Georgia
- PMID: 37901812
- PMCID: PMC10602647
- DOI: 10.3389/fmicb.2023.1258810
Metagenomics analysis reveals presence of the Merida-like virus in Georgia
Abstract
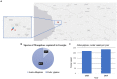
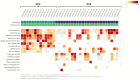
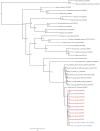
Arbovirus surveillance is fundamental for the discovery of novel viruses and prevention of febrile vector-borne illnesses. Vector-borne pathogens can rapidly expand and adapt in new geographic and environmental conditions. In this study, metagenomic surveillance was conducted to identify novel viruses in the Country of Georgia. A total of 521 mosquitoes were captured near a military training facility and pooled from species Culex pipiens (Linnaeus) (87%) and Aedes albopictus (Skuse) (13%). We decided to further analyze the Culex pipiens mosquitoes, due to the more extensive number of samples collected. Our approach was to utilize an unbiased total RNA-seq for pathogen discovery in order to explore the mosquito virome. The viral reads from this analysis were mostly aligned to Insect-specific viruses from two main families, the Iflaviridae; a positive-stranded RNA virus and the Rhabdoviridae; a negative- and single-stranded RNA virus. Our pathogen discovery analysis revealed viral reads aligning to the Merida-like virus Turkey (MERDLVT) strain among the Rhabdoviridae. To further validate this result, we conducted a BLAST sequence comparison analysis of our samples with the MERDLVT strain. Our positive samples aligned to the MERDLVT strain with 96-100% sequence identity and 99.7-100% sequence coverage. A bootstrapped maximum-likelihood phylogenetic tree was used to evaluate the evolutionary relationships among these positive pooled specimens with the (MERDLVT) strain. The Georgia samples clustered most closely with two strains from Turkey, the Merida-like virus KE-2017a isolate 139-1-21 and the Merida-like virus Turkey isolate P431. Collectively, these results show the presence of the MERDLVT strain in Georgia.
Keywords: Culex pipiens; Merida like-virus; NGS; mosquito; pathogen discovery; vector control.
Copyright © 2023 Potter-Birriel, Pollio, Knott, Chunashvili, Fung, Conte, Reinbold-Wasson and Hang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures

Similar articles
-
West Nile virus, Anopheles flavivirus, a novel flavivirus as well as Merida-like rhabdovirus Turkey in field-collected mosquitoes from Thrace and Anatolia.Infect Genet Evol. 2018 Jan;57:36-45. doi: 10.1016/j.meegid.2017.11.003. Epub 2017 Nov 8. Infect Genet Evol. 2018. PMID: 29128516
-
Metagenomic Analysis of the Virome of Mosquito Excreta.mSphere. 2020 Sep 9;5(5):e00587-20. doi: 10.1128/mSphere.00587-20. mSphere. 2020. PMID: 32907949 Free PMC article.
-
A novel rhabdovirus, related to Merida virus, in field-collected mosquitoes from Anatolia and Thrace.Arch Virol. 2017 Jul;162(7):1903-1911. doi: 10.1007/s00705-017-3314-4. Epub 2017 Mar 10. Arch Virol. 2017. PMID: 28283817
-
Small RNA responses of Culex mosquitoes and cell lines during acute and persistent virus infection.Insect Biochem Mol Biol. 2019 Jun;109:13-23. doi: 10.1016/j.ibmb.2019.04.008. Epub 2019 Apr 5. Insect Biochem Mol Biol. 2019. PMID: 30959110 Free PMC article.
-
Deciphering the Virome of Culex vishnui Subgroup Mosquitoes, the Major Vectors of Japanese Encephalitis, in Japan.Viruses. 2020 Feb 28;12(3):264. doi: 10.3390/v12030264. Viruses. 2020. PMID: 32121094 Free PMC article.
Cited by
-
Metagenome analysis of viruses associated with Anopheles mosquitoes from Ramu Upazila, Cox's Bazar District, Bangladesh.PeerJ. 2025 Mar 31;13:e19180. doi: 10.7717/peerj.19180. eCollection 2025. PeerJ. 2025. PMID: 40183042 Free PMC article.
-
Negevirus Piura Suppresses Zika Virus Replication in Mosquito Cells.Viruses. 2024 Feb 24;16(3):350. doi: 10.3390/v16030350. Viruses. 2024. PMID: 38543716 Free PMC article.
-
Characterization of the Virome in Mosquitoes Across Distinct Habitats in the Yucatán Peninsula, Mexico.Viruses. 2025 May 26;17(6):758. doi: 10.3390/v17060758. Viruses. 2025. PMID: 40573349 Free PMC article.
References
-
- Bolling B. G., Olea-Popelka F. J., Eisen L., Moore C. G., Blair C. D. (2012). Transmission dynamics of an insect-specific flavivirus in a naturally infected Culex pipiens laboratory colony and effects of co-infection on vector competence for West Nile virus. Virology 427, 90–97. doi: 10.1016/j.virol.2012.02.016 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

