Motifs in SARS-CoV-2 evolution
- PMID: 37903545
- PMCID: PMC10726165
- DOI: 10.1261/rna.079557.122
Motifs in SARS-CoV-2 evolution
Abstract
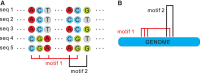
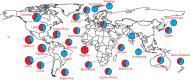
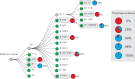
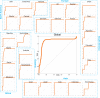
We present a novel framework enhancing the prediction of whether novel lineage poses the threat of eventually dominating the viral population. The framework is based purely on genomic sequence data, without requiring prior established biological analysis. Its building blocks are sets of coevolving sites in the alignment (motifs), identified via coevolutionary signals. The collection of such motifs forms a relational structure over the polymorphic sites. Motifs are constructed using distances quantifying the coevolutionary coupling of pairs and manifest as coevolving clusters of sites. We present an approach to genomic surveillance based on this notion of relational structure. Our system will issue an alert regarding a lineage, based on its contribution to drastic changes in the relational structure. We then conduct a comprehensive retrospective analysis of the COVID-19 pandemic based on SARS-CoV-2 genomic sequence data in GISAID from October 2020 to September 2022, across 21 lineages and 27 countries with weekly resolution. We investigate the performance of this surveillance system in terms of its accuracy, timeliness, and robustness. Lastly, we study how well each lineage is classified by such a system.
Keywords: SARS-CoV-2; coevolution; genomic surveillance; relational structure; site motif.
© 2024 Barrett et al.; Published by Cold Spring Harbor Laboratory Press for the RNA Society.
Figures



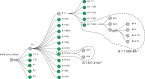
References
-
- Bacterial and Viral Bioinformatics Resource Center. 2022. Bacterial and viral bioinformatics resource center. https://www.bv-brc.org/. Last accessed Dec. 15, 2022.
-
- Barrett CL, Huang FW, Li TJ, Warren AS, Reidys CM. 2022. Rapid threat detection in SARS-CoV-2. medRxiv 10.1101/2022.08.05.22278480 - DOI
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous
