Clathrin light chains CLCa and CLCb have non-redundant roles in epithelial lumen formation
- PMID: 37923360
- PMCID: PMC10624596
- DOI: 10.26508/lsa.202302175
Clathrin light chains CLCa and CLCb have non-redundant roles in epithelial lumen formation
Abstract
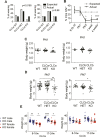
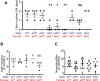
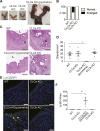
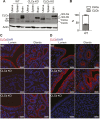
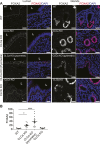
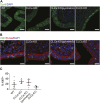
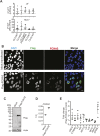
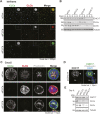
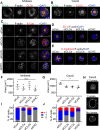
To identify functional differences between vertebrate clathrin light chains (CLCa or CLCb), phenotypes of mice lacking genes encoding either isoform were characterised. Mice without CLCa displayed 50% neonatal mortality, reduced body weight, reduced fertility, and ∼40% of aged females developed uterine pyometra. Mice lacking CLCb displayed a less severe weight reduction phenotype compared with those lacking CLCa and had no survival or reproductive system defects. Analysis of female mice lacking CLCa that developed pyometra revealed ectopic expression of epithelial differentiation markers (FOXA2 and K14) and a reduced number of endometrial glands, indicating defects in the lumenal epithelium. Defects in lumen formation and polarity of epithelial cysts derived from uterine or gut cell lines were also observed when either CLCa or CLCb were depleted, with more severe effects from CLCa depletion. In cysts, the CLC isoforms had different distributions relative to each other, although they converge in tissue. Together, these findings suggest differential and cooperative roles for CLC isoforms in epithelial lumen formation, with a dominant function for CLCa.
© 2023 Chen et al.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures




Similar articles
-
A unique role for clathrin light chain A in cell spreading and migration.J Cell Sci. 2019 May 15;132(10):jcs224030. doi: 10.1242/jcs.224030. J Cell Sci. 2019. PMID: 30975920 Free PMC article.
-
Clathrin light chain A drives selective myosin VI recruitment to clathrin-coated pits under membrane tension.Nat Commun. 2019 Oct 31;10(1):4974. doi: 10.1038/s41467-019-12855-6. Nat Commun. 2019. PMID: 31672988 Free PMC article.
-
Crosstalk between CLCb/Dyn1-Mediated Adaptive Clathrin-Mediated Endocytosis and Epidermal Growth Factor Receptor Signaling Increases Metastasis.Dev Cell. 2017 Feb 6;40(3):278-288.e5. doi: 10.1016/j.devcel.2017.01.007. Dev Cell. 2017. PMID: 28171750 Free PMC article.
-
Structure and function of CLCA proteins.Physiol Rev. 2005 Jul;85(3):1061-92. doi: 10.1152/physrev.00016.2004. Physiol Rev. 2005. PMID: 15987802 Review.
-
Role of the vascular endothelial growth factor isoforms in retinal angiogenesis and DiGeorge syndrome.Verh K Acad Geneeskd Belg. 2005;67(4):229-76. Verh K Acad Geneeskd Belg. 2005. PMID: 16334858 Review.
Cited by
-
CLCa mediates a novel cross-talk between Wnt secretion and actin organization.Life Sci Alliance. 2025 May 2;8(7):e202402962. doi: 10.26508/lsa.202402962. Print 2025 Jul. Life Sci Alliance. 2025. PMID: 40316417 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
