Applications of discriminative and deep learning feature extraction methods for whole slide image analysis: A survey
- PMID: 37928897
- PMCID: PMC10622844
- DOI: 10.1016/j.jpi.2023.100335
Applications of discriminative and deep learning feature extraction methods for whole slide image analysis: A survey
Abstract
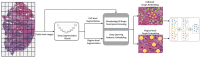
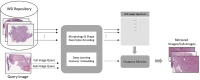
Digital pathology technologies, including whole slide imaging (WSI), have significantly improved modern clinical practices by facilitating storing, viewing, processing, and sharing digital scans of tissue glass slides. Researchers have proposed various artificial intelligence (AI) solutions for digital pathology applications, such as automated image analysis, to extract diagnostic information from WSI for improving pathology productivity, accuracy, and reproducibility. Feature extraction methods play a crucial role in transforming raw image data into meaningful representations for analysis, facilitating the characterization of tissue structures, cellular properties, and pathological patterns. These features have diverse applications in several digital pathology applications, such as cancer prognosis and diagnosis. Deep learning-based feature extraction methods have emerged as a promising approach to accurately represent WSI contents and have demonstrated superior performance in histology-related tasks. In this survey, we provide a comprehensive overview of feature extraction methods, including both manual and deep learning-based techniques, for the analysis of WSIs. We review relevant literature, analyze the discriminative and geometric features of WSIs (i.e., features suited to support the diagnostic process and extracted by "engineered" methods as opposed to AI), and explore predictive modeling techniques using AI and deep learning. This survey examines the advances, challenges, and opportunities in this rapidly evolving field, emphasizing the potential for accurate diagnosis, prognosis, and decision-making in digital pathology.
Keywords: Deep learning; Digital pathology applications; Image features engineering; Whole slide images.
Crown Copyright © 2023 Published by Elsevier Inc. on behalf of Association for Pathology Informatics.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures



Similar articles
-
Whole Slide Imaging, Artificial Intelligence, and Machine Learning in Pediatric and Perinatal Pathology: Current Status and Future Directions.Pediatr Dev Pathol. 2025 Mar-Apr;28(2):91-98. doi: 10.1177/10935266241299073. Epub 2024 Nov 18. Pediatr Dev Pathol. 2025. PMID: 39552500 Review.
-
Multilayer outperforms single-layer slide scanning in AI-based classification of whole slide images with low-burden acid-fast mycobacteria (AFB).Comput Methods Programs Biomed. 2023 Jun;234:107518. doi: 10.1016/j.cmpb.2023.107518. Epub 2023 Mar 28. Comput Methods Programs Biomed. 2023. PMID: 37018884
-
Enhancing Whole Slide Image Classification with Discriminative and Contrastive Learning.Med Image Comput Comput Assist Interv. 2024 Oct;15004:102-112. doi: 10.1007/978-3-031-72083-3_10. Epub 2024 Oct 14. Med Image Comput Comput Assist Interv. 2024. PMID: 40046787
-
Automated curation of large-scale cancer histopathology image datasets using deep learning.Histopathology. 2024 Jun;84(7):1139-1153. doi: 10.1111/his.15159. Epub 2024 Feb 26. Histopathology. 2024. PMID: 38409878
-
Artificial Intelligence in Lung Cancer Pathology Image Analysis.Cancers (Basel). 2019 Oct 28;11(11):1673. doi: 10.3390/cancers11111673. Cancers (Basel). 2019. PMID: 31661863 Free PMC article. Review.
Cited by
-
The role of nanomedicine and artificial intelligence in cancer health care: individual applications and emerging integrations-a narrative review.Discov Oncol. 2025 May 8;16(1):697. doi: 10.1007/s12672-025-02469-4. Discov Oncol. 2025. PMID: 40338421 Free PMC article. Review.
-
Advanced Deep Learning Approaches in Detection Technologies for Comprehensive Breast Cancer Assessment Based on WSIs: A Systematic Literature Review.Diagnostics (Basel). 2025 Apr 30;15(9):1150. doi: 10.3390/diagnostics15091150. Diagnostics (Basel). 2025. PMID: 40361968 Free PMC article. Review.
-
Advancing open-source visual analytics in digital pathology: A systematic review of tools, trends, and clinical applications.J Pathol Inform. 2025 May 23;18:100454. doi: 10.1016/j.jpi.2025.100454. eCollection 2025 Aug. J Pathol Inform. 2025. PMID: 40599690 Free PMC article. Review.
-
HcGAN: Harmonic conditional generative adversarial network for efficiently generating high-quality IHC images from H&E.Heliyon. 2024 Oct 1;10(20):e37902. doi: 10.1016/j.heliyon.2024.e37902. eCollection 2024 Oct 30. Heliyon. 2024. PMID: 39678164 Free PMC article.
-
An Unsupervised Learning Tool for Plaque Tissue Characterization in Histopathological Images.Sensors (Basel). 2024 Aug 20;24(16):5383. doi: 10.3390/s24165383. Sensors (Basel). 2024. PMID: 39205077 Free PMC article.
References
-
- Agus M., Al-Thelaya K., Calí C., et al. Eurographics Workshop on Visual Computing for Biology and Medicine. The Eurographics Association; 2020. InShaDe: invariant shape descriptors for visual analysis of histology 2D cellular and nuclear shapes; pp. 61–70. - DOI
-
- Ahmedt-Aristizabal D., Armin M.A., Denman S., Fookes C., Petersson L. A survey on graph-based deep learning for computational histopathology. Comput Med Imaging Graphics. 2021:102027. doi: 10.1016/j.compmedimag.2021.102027. https://www.sciencedirect.com/science/article/pii/S0895611121001762 - DOI - PubMed
Publication types
LinkOut - more resources
Full Text Sources

