ETV4-Dependent Transcriptional Plasticity Maintains MYC Expression and Results in IMiD Resistance in Multiple Myeloma
- PMID: 37934799
- PMCID: PMC10772538
- DOI: 10.1158/2643-3230.BCD-23-0061
ETV4-Dependent Transcriptional Plasticity Maintains MYC Expression and Results in IMiD Resistance in Multiple Myeloma
Abstract
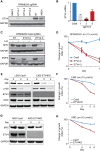
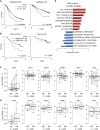
Immunomodulatory drugs (IMiD) are a backbone therapy for multiple myeloma (MM). Despite their efficacy, most patients develop resistance, and the mechanisms are not fully defined. Here, we show that IMiD responses are directed by IMiD-dependent degradation of IKZF1 and IKZF3 that bind to enhancers necessary to sustain the expression of MYC and other myeloma oncogenes. IMiD treatment universally depleted chromatin-bound IKZF1, but eviction of P300 and BRD4 coactivators only occurred in IMiD-sensitive cells. IKZF1-bound enhancers overlapped other transcription factor binding motifs, including ETV4. Chromatin immunoprecipitation sequencing showed that ETV4 bound to the same enhancers as IKZF1, and ETV4 CRISPR/Cas9-mediated ablation resulted in sensitization of IMiD-resistant MM. ETV4 expression is associated with IMiD resistance in cell lines, poor prognosis in patients, and is upregulated at relapse. These data indicate that ETV4 alleviates IKZF1 and IKZF3 dependency in MM by maintaining oncogenic enhancer activity and identify transcriptional plasticity as a previously unrecognized mechanism of IMiD resistance.
Significance: We show that IKZF1-bound enhancers are critical for IMiD efficacy and that the factor ETV4 can bind the same enhancers and substitute for IKZF1 and mediate IMiD resistance by maintaining MYC and other oncogenes. These data implicate transcription factor redundancy as a previously unrecognized mode of IMiD resistance in MM. See related article by Welsh, Barwick, et al., p. 34. See related commentary by Yun and Cleveland, p. 5. This article is featured in Selected Articles from This Issue, p. 4.
©2023 The Authors; Published by the American Association for Cancer Research.
Figures





Comment in
-
Transcriptional Plasticity Drives IMiD and p300 Inhibitor Resistance in Multiple Myeloma.Blood Cancer Discov. 2024 Jan 8;5(1):5-7. doi: 10.1158/2643-3230.BCD-23-0223. Blood Cancer Discov. 2024. PMID: 38085608 Free PMC article.
References
-
- Singhal S, Mehta J, Desikan R, Ayers D, Roberson P, Eddlemon P, et al. Antitumor activity of thalidomide in refractory multiple myeloma. N Engl J Med 1999;341:1565–71. - PubMed
-
- Raza S, Safyan RA, Lentzsch S. Immunomodulatory drugs (IMiDs) in multiple myeloma. Curr Cancer Drug Targets 2017;17:846–57. - PubMed
-
- Dimopoulos M, Spencer A, Attal M, Prince HM, Harousseau J-L, Dmoszynska A, et al. Lenalidomide plus dexamethasone for relapsed or refractory multiple myeloma. N Engl J Med 2007;357:2123–32. - PubMed
-
- Weber DM, Chen C, Niesvizky R, Wang M, Belch A, Stadtmauer EA, et al. Lenalidomide plus dexamethasone for relapsed multiple myeloma in North America. N Engl J Med 2007;357:2133–42. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

